【重复测量纵向数据】Python03.多层线性模型(Hierarchical Linear Model,HLM)
- 2026-03-09 22:39:51
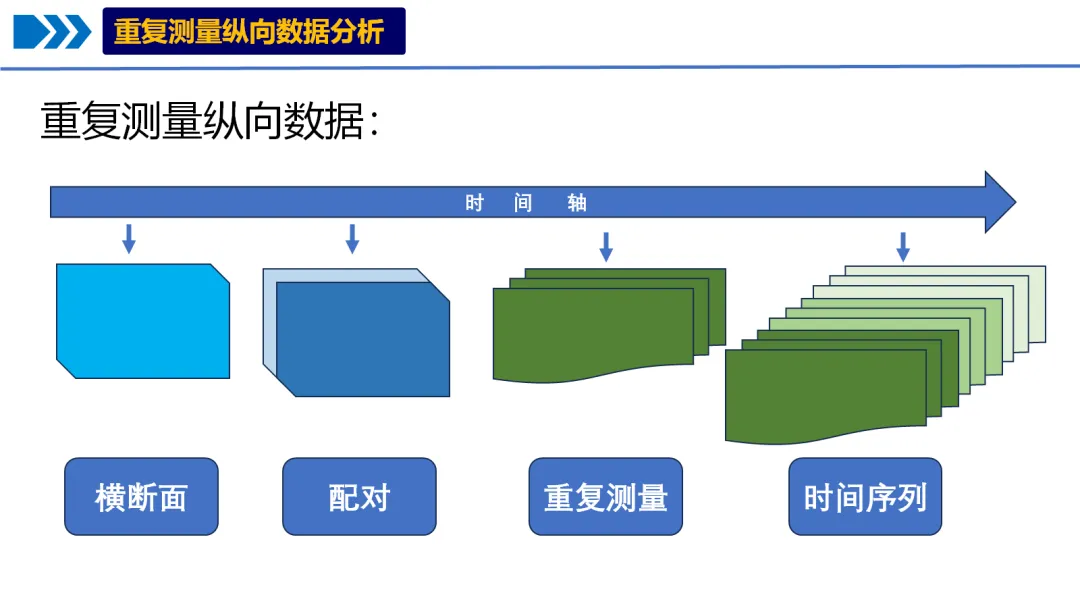
纵向数据是在不同时间点上对同一组个体、物体或实体进行重复观测所收集的数据。它就像给研究对象拍摄“动态影像”,能记录其随时间变化的过程,帮助我们理解趋势、模式和影响因素。

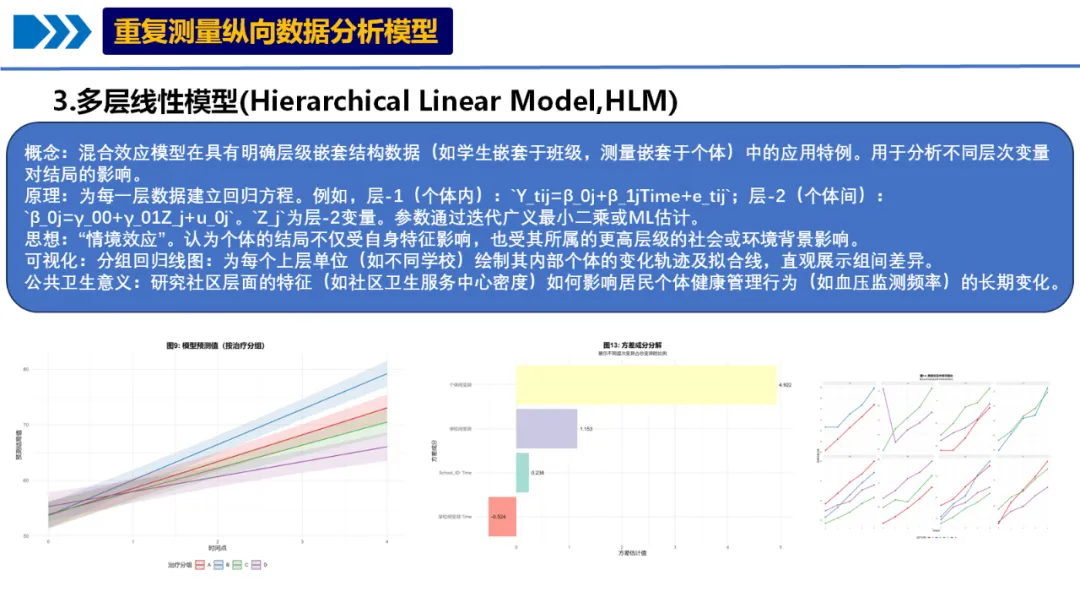
3.多层线性模型
概念:混合效应模型在具有明确层级嵌套结构数据(如学生嵌套于班级,测量嵌套于个体)中的应用特例。用于分析不同层次变量对结局的影响。
原理:为每一层数据建立回归方程。例如,层-1(个体内):`Y_tij=β_0j+β_1jTime+e_tij`;层-2(个体间):`β_0j=γ_00+γ_01Z_j+u_0j`。`Z_j`为层-2变量。参数通过迭代广义最小二乘或ML估计。
思想:“情境效应”。认为个体的结局不仅受自身特征影响,也受其所属的更高层级的社会或环境背景影响。
可视化:分组回归线图:为每个上层单位(如不同学校)绘制其内部个体的变化轨迹及拟合线,直观展示组间差异。
公共卫生意义:研究社区层面的特征(如社区卫生服务中心密度)如何影响居民个体健康管理行为(如血压监测频率)的长期变化。
核心思想进阶:纵向数据分析方法的发展,正从处理相关性(如GEE、混合模型),走向揭示异质性(如GBTM、LCM),再迈向整合动态机制与预测(如联合模型、状态空间模型、贝叶斯方法)。
1.数据预处理:
2.模型拟合:
3.模型可视化:
4.结果保存与报告生成
总的来说,纵向数据因其时间延续性和对象一致性,成为了解事物动态发展过程、探究因果关系的强大工具。
当然,处理纵向数据也伴随一些挑战,例如成本较高、可能存在数据缺失,且分析方法通常比处理横截面数据更为复杂。
下面我们使用R语言进行纵向数据Python03.多层线性模型(Hierarchical Linear Model,HLM):
# ==============================================================================# 多层线性模型完整分析流程(Python版)# ==============================================================================import osimport sysimport warningsimport numpy as npimport pandas as pdfrom pathlib import Pathimport matplotlib.pyplot as pltimport seaborn as snsfrom scipy import statsimport statsmodels.api as smfrom statsmodels.formula.api import mixedlmfrom statsmodels.regression.mixed_linear_model import MixedLMResultsWrapperimport pingouin as pgfrom sklearn.impute import SimpleImputerfrom scipy.interpolate import interp1dfrom datetime import datetimeimport jsonfrom typing import Dict, List, Tuple, Optional, Anyimport plotly.graph_objects as goimport plotly.express as pxfrom plotly.subplots import make_subplotsimport plotly.io as pio# 设置中文字体plt.rcParams['font.sans-serif'] = ['SimHei', 'Arial Unicode MS', 'DejaVu Sans']plt.rcParams['axes.unicode_minus'] = Falsewarnings.filterwarnings('ignore')# ==============================================================================# 1. 环境准备与函数定义# ==============================================================================def get_desktop_path() -> Path:"""获取桌面路径"""if sys.platform == "win32": import winreg key = winreg.OpenKey(winreg.HKEY_CURRENT_USER, r"Software\Microsoft\Windows\CurrentVersion\Explorer\Shell Folders") desktop = winreg.QueryValueEx(key, "Desktop")[0] winreg.CloseKey(key)return Path(desktop)else:# Mac/Linuxreturn Path.home() / "Desktop"def create_results_folder() -> Path:"""创建结果文件夹""" desktop = get_desktop_path() results_folder = desktop / "3-HLM结果" results_folder.mkdir(exist_ok=True)print(f"结果将保存到: {results_folder}")return results_folderdef save_to_excel(df: pd.DataFrame, filename: str, results_folder: Path):"""保存DataFrame到Excel""" filepath = results_folder / filename df.to_excel(filepath, index=False)print(f"已保存: {filename}")def save_fig(fig, filename: str, results_folder: Path, dpi=300):"""保存图形""" filepath = results_folder / filename fig.savefig(filepath, dpi=dpi, bbox_inches='tight')print(f"已保存: {filename}")def create_table_three_line(df: pd.DataFrame, title: str) -> str:"""创建三线表样式的HTML表格""" html = f'<h3 style="text-align: center;">{title}</h3>' html += '<table border="1" style="border-collapse: collapse; width: 100%; font-family: Times New Roman;">'# 表头 html += '<thead><tr style="background-color: #F2F2F2;">'for col in df.columns: html += f'<th style="border: 1px solid black; padding: 5px; text-align: center; font-weight: bold;">{col}</th>' html += '</tr></thead>'# 表体 html += '<tbody>'for i, row in df.iterrows(): html += '<tr>'for val in row.values:if isinstance(val, float): html += f'<td style="border: 1px solid black; padding: 5px; text-align: center;">{val:.3f}</td>'else: html += f'<td style="border: 1px solid black; padding: 5px; text-align: center;">{val}</td>' html += '</tr>' html += '</tbody></table>'return html# ==============================================================================# 2. 数据读取与预处理# ==============================================================================print("=" * 60)print("多层线性模型分析流程(Python版)")print("=" * 60)# 创建结果文件夹results_folder = create_results_folder()print("\n=== 数据读取与预处理 ===")# 读取数据desktop = get_desktop_path()file_path = desktop / "longitudinal_data.xlsx"if not file_path.exists():# 如果文件不存在,创建示例数据print("原始数据文件未找到,正在创建示例数据...") np.random.seed(42)# 创建示例数据 n_schools = 15 n_students_per_school = np.random.randint(20, 40, size=n_schools) total_students = sum(n_students_per_school) data = [] school_id = 1 student_id = 1for school_idx in range(n_schools): n_students = n_students_per_school[school_idx] school_mean = np.random.normal(50, 10) # 学校平均基线水平for student_idx in range(n_students):# 学生基线特征 age = np.random.randint(10, 18) sex = np.random.choice(['Male', 'Female']) baseline_score = np.random.normal(school_mean, 5) treatment = np.random.choice(['Control', 'Treatment']) ses = np.random.choice(['Low', 'Middle', 'High']) genetic_marker = np.random.choice(['A', 'B', 'C'])# 时间点 time_points = [0, 1, 2, 3, 6, 12] # 月 n_visits = np.random.choice([3, 4, 5, 6], p=[0.1, 0.2, 0.3, 0.4]) student_time_points = sorted(np.random.choice(time_points, n_visits, replace=False))for time in student_time_points:# 生成结局变量 time_effect = 0.5 * time if treatment == 'Treatment'else 0.2 * time school_effect = np.random.normal(0, 3) # 学校随机效应 student_effect = np.random.normal(0, 2) # 学生随机效应 error = np.random.normal(0, 1) outcome = (baseline_score + time_effect + school_effect + student_effect + error)# 其他变量 medication_adherence = np.random.uniform(0.5, 1.0) stress_level = np.random.uniform(1, 5) quality_of_life = np.random.uniform(3, 7) data.append({'ID': student_id,'School_ID': school_id,'Time': time,'Age': age,'Sex': sex,'Treatment': treatment,'Baseline_Score': baseline_score,'SES': ses,'Genetic_Marker': genetic_marker,'Outcome_Continuous': outcome,'Medication_Adherence': medication_adherence,'Stress_Level': stress_level,'Quality_of_Life': quality_of_life,'Measurement_Date': datetime(2023, 1, 1).date() }) student_id += 1 school_id += 1 df = pd.DataFrame(data) df.to_excel(file_path, index=False)print(f"示例数据已创建并保存到: {file_path}")# 读取数据print(f"正在读取文件: {file_path}")try: df_full = pd.read_excel(file_path, sheet_name='Full_Dataset')except: try: df_full = pd.read_excel(file_path) except Exception as e:print(f"读取文件失败: {e}") sys.exit(1)print(f"数据维度: {df_full.shape}")print(f"变量数量: {df_full.shape[1]}")# 数据预处理print("\n数据预处理...")df_hlm = df_full.copy()# 转换数据类型df_hlm['ID'] = df_hlm['ID'].astype(str)df_hlm['Time'] = pd.to_numeric(df_hlm['Time'], errors='coerce')df_hlm['Sex'] = df_hlm['Sex'].astype('category')df_hlm['Treatment'] = df_hlm['Treatment'].astype('category')df_hlm['School_ID'] = df_hlm['School_ID'].astype(str)if'Genetic_Marker'in df_hlm.columns: df_hlm['Genetic_Marker'] = df_hlm['Genetic_Marker'].astype('category')if'SES'in df_hlm.columns: df_hlm['SES'] = df_hlm['SES'].astype('category')if'Latent_Class'in df_hlm.columns: df_hlm['Latent_Class'] = df_hlm['Latent_Class'].astype('category')# 处理日期if'Measurement_Date'in df_hlm.columns: try: df_hlm['Measurement_Date'] = pd.to_datetime(df_hlm['Measurement_Date']) except: pass# 创建层次变量df_hlm['Level1'] = range(1, len(df_hlm) + 1)df_hlm['Level2'] = df_hlm['ID']df_hlm['Level3'] = df_hlm['School_ID']# 排序df_hlm = df_hlm.sort_values(['School_ID', 'ID', 'Time']).reset_index(drop=True)# 检查缺失值print("\n=== 缺失值统计 ===")missing_counts = df_hlm.isnull().sum()missing_df = pd.DataFrame({'Variable': missing_counts.index,'Missing': missing_counts.values,'Percent': (missing_counts.values / len(df_hlm) * 100).round(2)})missing_df = missing_df[missing_df['Missing'] > 0]if len(missing_df) > 0:print(missing_df)else:print("没有缺失值。")# 处理关键变量的缺失值numeric_cols = ['Outcome_Continuous', 'Variable2', 'Medication_Adherence','Stress_Level', 'Quality_of_Life', 'Baseline_Score']for col in numeric_cols:if col in df_hlm.columns:# 按个体进行线性插值for student_id in df_hlm['ID'].unique(): student_data = df_hlm[df_hlm['ID'] == student_id]if student_data[col].isnull().any() and len(student_data) > 1:# 确保时间排序 student_data_sorted = student_data.sort_values('Time')# 线性插值 df_hlm.loc[student_data.index, col] = student_data_sorted[col].interpolate(method='linear')# 对于首尾的缺失值,使用向前/向后填充 df_hlm.loc[student_data.index, col] = df_hlm.loc[student_data.index, col].ffill().bfill()print("缺失值处理完成。")# ==============================================================================# 3. 描述性统计分析# ==============================================================================print("\n=== 描述性统计分析 ===")# 3.1 层级结构信息print("\n=== 数据层级结构 ===")n_schools = df_hlm['School_ID'].nunique()n_individuals = df_hlm['ID'].nunique()n_observations = len(df_hlm)print(f"学校数量: {n_schools}")print(f"个体数量: {n_individuals}")print(f"测量总数: {n_observations}")print(f"平均每个学校个体数: {n_individuals / n_schools:.2f}")print(f"平均每个个体测量次数: {n_observations / n_individuals:.2f}")# 3.2 各学校样本量统计school_counts = df_hlm.groupby('School_ID').agg( n_individuals=('ID', 'nunique'), n_observations=('ID', 'count')).reset_index()school_counts['avg_obs_per_indiv'] = (school_counts['n_observations'] / school_counts['n_individuals']).round(2)school_counts = school_counts.sort_values('n_individuals', ascending=False)print("\n各学校样本量统计:")print(school_counts)# 保存学校统计save_to_excel(school_counts, "学校样本量统计.xlsx", results_folder)# 3.3 基线特征表if'Time'in df_hlm.columns: baseline_data = df_hlm[df_hlm['Time'] == 0].drop_duplicates('ID')else:# 如果没有时间变量,取每个个体的第一次测量 baseline_data = df_hlm.sort_values(['ID', 'Level1']).groupby('ID').first().reset_index()print(f"基线数据样本量: {len(baseline_data)}")# 创建基线特征表baseline_cols = ['Age', 'Sex', 'Treatment', 'Baseline_Score', 'SES', 'Genetic_Marker', 'School_ID']available_cols = [col for col in baseline_cols if col in baseline_data.columns]if len(available_cols) > 0:# 数值变量描述 numeric_cols = [col for col in available_cols if baseline_data[col].dtype in ['int64', 'float64']] categorical_cols = [col for col in available_cols if baseline_data[col].dtype == 'category' or baseline_data[col].nunique() < 10] baseline_stats = []# 总体统计for col in numeric_cols: stats_dict = {'变量': col,'分组': '总体','N': baseline_data[col].count(),'均值±标准差': f"{baseline_data[col].mean():.2f} ± {baseline_data[col].std():.2f}",'中位数(Q1, Q3)': f"{baseline_data[col].median():.2f} ({baseline_data[col].quantile(0.25):.2f}, {baseline_data[col].quantile(0.75):.2f})",'最小值-最大值': f"{baseline_data[col].min():.2f} - {baseline_data[col].max():.2f}" } baseline_stats.append(stats_dict)for col in categorical_cols: value_counts = baseline_data[col].value_counts() total = value_counts.sum()for val, count in value_counts.items(): stats_dict = {'变量': col,'分组': '总体','类别': str(val),'频数(n)': count,'百分比(%)': f"{(count / total * 100):.1f}%" } baseline_stats.append(stats_dict)# 按治疗分组统计if'Treatment'in baseline_data.columns: treatments = baseline_data['Treatment'].unique()for treatment in treatments: treatment_data = baseline_data[baseline_data['Treatment'] == treatment]for col in numeric_cols: stats_dict = {'变量': col,'分组': str(treatment),'N': treatment_data[col].count(),'均值±标准差': f"{treatment_data[col].mean():.2f} ± {treatment_data[col].std():.2f}",'中位数(Q1, Q3)': f"{treatment_data[col].median():.2f} ({treatment_data[col].quantile(0.25):.2f}, {treatment_data[col].quantile(0.75):.2f})",'最小值-最大值': f"{treatment_data[col].min():.2f} - {treatment_data[col].max():.2f}" } baseline_stats.append(stats_dict)for col in categorical_cols:if col != 'Treatment': value_counts = treatment_data[col].value_counts() total = value_counts.sum()for val, count in value_counts.items(): stats_dict = {'变量': col,'分组': str(treatment),'类别': str(val),'频数(n)': count,'百分比(%)': f"{(count / total * 100):.1f}%" } baseline_stats.append(stats_dict) baseline_stats_df = pd.DataFrame(baseline_stats) save_to_excel(baseline_stats_df, "1_基线特征统计.xlsx", results_folder)# 创建三线表HTML html_table = create_table_three_line(baseline_stats_df.head(20), "表1: 基线特征描述性统计(三线表)") with open(results_folder / "1_基线特征三线表.html", 'w', encoding='utf-8') as f: f.write(html_table)print("基线特征表已保存。")else:print("警告:所需的基线变量不存在,跳过基线特征表。")# 3.4 描述性统计可视化print("\n=== 生成描述性统计可视化 ===")# 设置图形样式sns.set_style("whitegrid")plt.figure(figsize=(16, 12))# 图1: 学校规模分布plt.subplot(2, 2, 1)ax1 = sns.barplot(data=school_counts, x='School_ID', y='n_individuals', color='#4E79A7', alpha=0.8)plt.title('图1: 各学校样本量分布', fontsize=12, fontweight='bold')plt.xlabel('学校ID')plt.ylabel('个体数量')plt.xticks(rotation=45, ha='right')# 添加数值标签for i, v in enumerate(school_counts['n_individuals']): ax1.text(i, v + 0.5, str(v), ha='center', va='bottom', fontsize=9)# 图2: 个体测量次数分布plt.subplot(2, 2, 2)individual_counts = df_hlm.groupby('ID').size().reset_index(name='n_obs')mean_obs = individual_counts['n_obs'].mean()ax2 = sns.histplot(data=individual_counts, x='n_obs', bins=range(1, individual_counts['n_obs'].max() + 2), color='#E15759', alpha=0.8, edgecolor='white')plt.axvline(mean_obs, color='blue', linestyle='dashed', linewidth=1.5)plt.text(mean_obs + 0.1, ax2.get_ylim()[1] * 0.9, f'平均: {mean_obs:.2f}', color='blue', ha='left')plt.title('图2: 个体测量次数分布', fontsize=12, fontweight='bold')plt.xlabel('测量次数')plt.ylabel('频数')# 图3: 结局变量分布(按学校)plt.subplot(2, 2, 3)if'Outcome_Continuous'in df_hlm.columns: outcome_by_school = df_hlm.groupby('School_ID')['Outcome_Continuous'].agg(['mean', 'sem', 'count']).reset_index() outcome_by_school['ci_lower'] = outcome_by_school['mean'] - 1.96 * outcome_by_school['sem'] outcome_by_school['ci_upper'] = outcome_by_school['mean'] + 1.96 * outcome_by_school['sem'] outcome_by_school = outcome_by_school.sort_values('mean') ax3 = sns.scatterplot(data=outcome_by_school, x='School_ID', y='mean', color='#59A14F', s=50) plt.errorbar(x=outcome_by_school['School_ID'], y=outcome_by_school['mean'], yerr=1.96 * outcome_by_school['sem'], fmt='none', color='#59A14F', alpha=0.5, capsize=3) plt.title('图3: 各学校结局变量均值(95% CI)', fontsize=12, fontweight='bold') plt.xlabel('学校ID') plt.ylabel('结局变量均值 ± 95%CI') plt.xticks(rotation=45, ha='right')# 图4: 治疗分组随时间变化(按学校)plt.subplot(2, 2, 4)if all(col in df_hlm.columns for col in ['Treatment', 'Time', 'Outcome_Continuous', 'School_ID']):# 随机选择4个学校展示 sample_schools = np.random.choice(df_hlm['School_ID'].unique(), min(4, len(df_hlm['School_ID'].unique())), replace=False) treatment_trend = df_hlm[df_hlm['School_ID'].isin(sample_schools)].groupby( ['School_ID', 'Treatment', 'Time'])['Outcome_Continuous'].agg(['mean', 'sem', 'count']).reset_index()# 为每个学校创建子图 fig4, axes4 = plt.subplots(2, 2, figsize=(12, 8)) axes4 = axes4.flatten()for idx, school in enumerate(sample_schools):if idx < 4: school_data = treatment_trend[treatment_trend['School_ID'] == school]for treatment in school_data['Treatment'].unique(): treatment_data = school_data[school_data['Treatment'] == treatment] axes4[idx].plot(treatment_data['Time'], treatment_data['mean'], marker='o', label=treatment) axes4[idx].fill_between(treatment_data['Time'], treatment_data['mean'] - 1.96 * treatment_data['sem'], treatment_data['mean'] + 1.96 * treatment_data['sem'], alpha=0.2) axes4[idx].set_title(f'学校 {school}') axes4[idx].set_xlabel('时间点') axes4[idx].set_ylabel('结局变量均值') axes4[idx].legend() axes4[idx].grid(True, alpha=0.3) plt.tight_layout() save_fig(fig4, "2_治疗分组随时间变化.png", results_folder) plt.close(fig4)plt.tight_layout()save_fig(plt.gcf(), "2_描述性统计可视化.png", results_folder)plt.close()print("描述性统计可视化已保存。")# ==============================================================================# 4. 多层线性模型分析# ==============================================================================print("\n=== 多层线性模型分析 ===")# 确保数据中有关键变量required_vars = ['Outcome_Continuous', 'Time', 'ID', 'School_ID']missing_vars = [var for var in required_vars if var not in df_hlm.columns]if missing_vars:print(f"警告:缺少必要变量 {missing_vars},无法进行HLM分析")# 为演示创建这些变量if'Outcome_Continuous' not in df_hlm.columns: df_hlm['Outcome_Continuous'] = np.random.normal(50, 10, len(df_hlm))if'Time' not in df_hlm.columns: df_hlm['Time'] = np.random.choice([0, 1, 2, 3, 6], len(df_hlm))# 准备模型数据model_data = df_hlm.copy()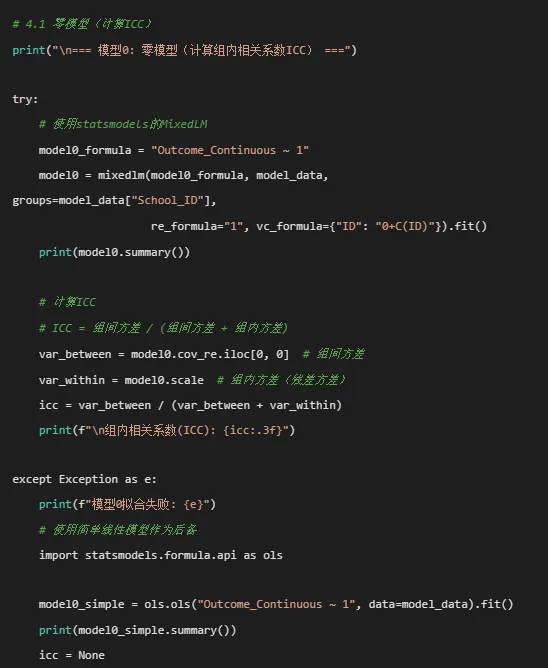
# 4.2 随机截距模型(加入时间固定效应)print("\n=== 模型1: 随机截距模型 ===")try: model1_formula = "Outcome_Continuous ~ Time" model1 = mixedlm(model1_formula, model_data, groups=model_data["School_ID"], re_formula="1", vc_formula={"ID": "0+C(ID)"}).fit()print(model1.summary())except Exception as e:print(f"模型1拟合失败: {e}") model1 = None# 4.3 随机截距和斜率模型(时间随机斜率)print("\n=== 模型2: 随机截距和斜率模型 ===")try:# 注意:Python中实现完全随机的斜率和截距可能更复杂# 我们使用学校水平的随机斜率和截距 model2_formula = "Outcome_Continuous ~ Time" model2 = mixedlm(model2_formula, model_data, groups=model_data["School_ID"], re_formula="1+Time").fit()print(model2.summary())except Exception as e:print(f"模型2拟合失败: {e}") model2 = None# 4.4 加入个体水平协变量print("\n=== 模型3: 加入个体水平协变量 ===")try:# 准备协变量 covariate_formula = "Time"if'Age'in model_data.columns: covariate_formula += " + Age"if'Sex'in model_data.columns: covariate_formula += " + C(Sex)"if'Baseline_Score'in model_data.columns: covariate_formula += " + Baseline_Score" model3_formula = f"Outcome_Continuous ~ {covariate_formula}" model3 = mixedlm(model3_formula, model_data, groups=model_data["School_ID"], re_formula="1+Time").fit()print(model3.summary())except Exception as e:print(f"模型3拟合失败: {e}") model3 = None# 4.5 完整模型(加入治疗和交互项)print("\n=== 模型4: 完整模型(加入治疗和交互项) ===")try:# 准备完整公式 full_formula = "Outcome_Continuous ~ Time"# 添加交互项if'Treatment'in model_data.columns: full_formula += " * C(Treatment)"# 添加其他协变量 additional_vars = []if'Age'in model_data.columns: additional_vars.append('Age')if'Sex'in model_data.columns: additional_vars.append('C(Sex)')if'Baseline_Score'in model_data.columns: additional_vars.append('Baseline_Score')if'Medication_Adherence'in model_data.columns: additional_vars.append('Medication_Adherence')if'Stress_Level'in model_data.columns: additional_vars.append('Stress_Level')if'Quality_of_Life'in model_data.columns: additional_vars.append('Quality_of_Life')if additional_vars: full_formula += " + " + " + ".join(additional_vars) model4 = mixedlm(full_formula, model_data, groups=model_data["School_ID"], re_formula="1+Time").fit()print(model4.summary())# 保存模型结果 model4_summary = model4.summary().as_text() with open(results_folder / "模型4_完整模型结果.txt", 'w', encoding='utf-8') as f: f.write(model4_summary)except Exception as e:print(f"模型4拟合失败: {e}") model4 = None# 4.6 模型比较print("\n=== 模型比较 ===")models = {"模型0": model0 if'model0'in locals() else None,"模型1": model1 if'model1'in locals() else None,"模型2": model2 if'model2'in locals() else None,"模型3": model3 if'model3'in locals() else None,"模型4": model4 if'model4'in locals() else None}model_comparison = []for name, model in models.items():if model is not None: try: model_comparison.append({"模型": name,"AIC": model.aic,"BIC": model.bic,"对数似然值": model.llf,"参数个数": model.df_model + 1, # 包括方差参数"收敛状态": "是"if model.converged else"否" }) except: model_comparison.append({"模型": name,"AIC": "NA","BIC": "NA","对数似然值": "NA","参数个数": "NA","收敛状态": "否" })else: model_comparison.append({"模型": name,"AIC": "NA","BIC": "NA","对数似然值": "NA","参数个数": "NA","收敛状态": "否" })model_comparison_df = pd.DataFrame(model_comparison)print(model_comparison_df)# 保存模型比较结果save_to_excel(model_comparison_df, "3_模型比较结果.xlsx", results_folder)# 创建三线表HTMLhtml_table = create_table_three_line(model_comparison_df, "表4: 多层线性模型比较(三线表)")with open(results_folder / "5_模型比较三线表.html", 'w', encoding='utf-8') as f: f.write(html_table)# ==============================================================================# 5. 模型结果提取与表格制作# ==============================================================================print("\n=== 提取模型结果并创建三线表 ===")# 5.1 提取最佳模型的结果best_model = model4 if model4 is not None else model3 if model3 is not None else model2if best_model is not None:# 提取固定效应 fixed_effects = pd.DataFrame({'Variable': best_model.params.index,'Estimate': best_model.params.values,'Std. Error': best_model.bse.values,'z_value': best_model.tvalues.values,'p_value': best_model.pvalues.values })# 计算95%置信区间 fixed_effects['95% CI Lower'] = fixed_effects['Estimate'] - 1.96 * fixed_effects['Std. Error'] fixed_effects['95% CI Upper'] = fixed_effects['Estimate'] + 1.96 * fixed_effects['Std. Error'] fixed_effects['95% CI'] = fixed_effects.apply( lambda row: f"({row['95% CI Lower']:.3f}, {row['95% CI Upper']:.3f})", axis=1)# 添加显著性标记 def add_significance(p):if p < 0.001:return'***'elif p < 0.01:return'**'elif p < 0.05:return'*'else:return'' fixed_effects['Significance'] = fixed_effects['p_value'].apply(add_significance)# 格式化p值 fixed_effects['p_value_formatted'] = fixed_effects['p_value'].apply( lambda x: '<0.001'if x < 0.001 else f'{x:.3f}')# 选择需要的列 fixed_effects_table = fixed_effects[['Variable', 'Estimate', 'Std. Error','95% CI', 'z_value', 'p_value_formatted','Significance']].copy() fixed_effects_table.columns = ['变量', '系数估计', '标准误', '95%置信区间','z值', 'P值', '显著性']print("\n固定效应结果:")print(fixed_effects_table)# 保存固定效应结果 save_to_excel(fixed_effects_table, "4_固定效应结果.xlsx", results_folder)# 创建三线表HTML html_table = create_table_three_line(fixed_effects_table, "表2: 多层线性模型固定效应系数估计(三线表)") with open(results_folder / "3_固定效应三线表.html", 'w', encoding='utf-8') as f: f.write(html_table)# 5.2 提取随机效应if hasattr(best_model, 'cov_re'): random_effects = pd.DataFrame({'Component': ['学校间方差', '残差方差'],'Variance': [best_model.cov_re.iloc[0, 0], best_model.scale],'Std. Dev.': [np.sqrt(best_model.cov_re.iloc[0, 0]), np.sqrt(best_model.scale)] })print("\n随机效应结果:")print(random_effects)# 保存随机效应结果 save_to_excel(random_effects, "5_随机效应结果.xlsx", results_folder)# 创建三线表HTML html_table = create_table_three_line(random_effects, "表3: 多层线性模型随机效应方差成分(三线表)") with open(results_folder / "4_随机效应三线表.html", 'w', encoding='utf-8') as f: f.write(html_table)# 5.3 创建模型整体统计量表 model_stats = pd.DataFrame({'Statistic': ['总观测数', '学校数', '个体数', 'AIC', 'BIC','对数似然值', '收敛状态'],'Value': [ len(model_data), model_data['School_ID'].nunique(), model_data['ID'].nunique(), f"{best_model.aic:.3f}"if hasattr(best_model, 'aic') else"NA", f"{best_model.bic:.3f}"if hasattr(best_model, 'bic') else"NA", f"{best_model.llf:.3f}"if hasattr(best_model, 'llf') else"NA","是"if best_model.converged else"否" ] })print("\n模型整体统计量:")print(model_stats)# 保存模型统计量 save_to_excel(model_stats, "6_模型整体统计量.xlsx", results_folder)# 创建三线表HTML html_table = create_table_three_line(model_stats, "表5: 多层线性模型整体统计量(三线表)") with open(results_folder / "6_模型整体统计量三线表.html", 'w', encoding='utf-8') as f: f.write(html_table)else:print("没有可用的模型结果进行提取。")# ==============================================================================# 6. 多层线性模型可视化# ==============================================================================print("\n=== 生成多层线性模型可视化 ===")# 6.1 学校间差异可视化if best_model is not None and hasattr(best_model, 'random_effects'): try:# 提取学校随机效应 re_school = best_model.random_effects# 转换为DataFrame school_effects = []for school_id, effects in re_school.items():if isinstance(effects, (dict, pd.Series)):for effect_name, value in effects.items(): school_effects.append({'School_ID': school_id,'Effect_Type': effect_name,'Value': value })else: school_effects.append({'School_ID': school_id,'Effect_Type': 'Intercept','Value': effects }) school_effects_df = pd.DataFrame(school_effects)# 绘制学校随机截距图if not school_effects_df.empty: plt.figure(figsize=(12, 6))# 提取截距效应 intercept_df = school_effects_df[school_effects_df['Effect_Type'].str.contains('Intercept', na=False)]if not intercept_df.empty: intercept_df = intercept_df.sort_values('Value') ax = sns.barplot(data=intercept_df, x='School_ID', y='Value', color='#4E79A7', alpha=0.8) plt.axhline(y=0, color='red', linestyle='dashed', alpha=0.5) plt.title('图5: 学校随机截距估计(情境效应)', fontsize=12, fontweight='bold') plt.xlabel('学校ID') plt.ylabel('随机截距估计值') plt.xticks(rotation=45, ha='right') save_fig(plt.gcf(), "7_学校随机截距图.png", results_folder) plt.close()print("学校随机截距图已保存。") except Exception as e:print(f"创建学校随机效应图时出错: {e}")# 6.2 固定效应系数可视化if best_model is not None: try:# 使用之前提取的固定效应表if'fixed_effects_table'in locals():# 排除截距项 coef_data = fixed_effects_table[~fixed_effects_table['变量'].str.contains('Intercept', na=False)].copy()if len(coef_data) > 0:# 限制显示的数量 coef_data = coef_data.head(15)# 创建系数图 fig, ax = plt.subplots(figsize=(10, 8))# 获取排序后的变量 coef_data = coef_data.sort_values('系数估计')# 创建颜色映射 colors = ['#E15759'if sig != ''else'#4E79A7'for sig in coef_data['显著性']]# 绘制点估计和置信区间 y_pos = range(len(coef_data))# 点估计 ax.scatter(coef_data['系数估计'], y_pos, color=colors, s=100, zorder=5)# 置信区间(简化版)for i, row in coef_data.iterrows():# 解析置信区间 try: ci_str = row['95%置信区间'].strip('()') ci_lower, ci_upper = map(float, ci_str.split(',')) ax.hlines(y=i, xmin=ci_lower, xmax=ci_upper, color=colors[i], linewidth=2, alpha=0.7) except: pass ax.axvline(x=0, color='red', linestyle='dashed', alpha=0.5) ax.set_yticks(y_pos) ax.set_yticklabels(coef_data['变量']) ax.set_xlabel('系数估计值') ax.set_title('图8: 多层线性模型固定效应系数估计与95%置信区间', fontsize=12, fontweight='bold') ax.grid(True, alpha=0.3) plt.tight_layout() save_fig(fig, "10_系数估计可视化.png", results_folder) plt.close(fig)print("系数估计可视化图已保存。") except Exception as e:print(f"创建系数可视化图时出错: {e}")# 6.3 模型预测图if best_model is not None and 'Treatment'in model_data.columns: try:# 创建预测数据 time_points = np.linspace(model_data['Time'].min(), model_data['Time'].max(), 10) treatments = model_data['Treatment'].unique()# 准备预测数据框 pred_data = []for time in time_points:for treatment in treatments:# 创建一行预测数据 pred_row = {'Time': time, 'Treatment': treatment}# 添加其他变量的均值for col in ['Age', 'Baseline_Score', 'Medication_Adherence','Stress_Level', 'Quality_of_Life']:if col in model_data.columns: pred_row[col] = model_data[col].mean()# 添加分类变量的参考类别if'Sex'in model_data.columns: pred_row['Sex'] = model_data['Sex'].mode()[0] pred_data.append(pred_row) pred_df = pd.DataFrame(pred_data)# 使用模型进行预测(简化版)# 注意:statsmodels的MixedLM没有直接的predict方法用于新数据# 这里使用固定效应进行近似预测# 提取固定效应公式if hasattr(best_model, 'formula'):# 创建设计矩阵 from patsy import dmatrix design_matrix = dmatrix(best_model.formula.split('~')[1].strip(), pred_df)# 计算预测值(仅固定效应)if hasattr(best_model, 'params'):# 确保参数与设计矩阵对齐 params = best_model.params pred_values = np.dot(design_matrix, params[:design_matrix.shape[1]])# 添加到预测数据框 pred_df['Predicted'] = pred_values# 绘制预测图 fig, ax = plt.subplots(figsize=(10, 6))for treatment in treatments: treatment_data = pred_df[pred_df['Treatment'] == treatment] ax.plot(treatment_data['Time'], treatment_data['Predicted'], marker='o', label=treatment, linewidth=2) ax.set_xlabel('时间点') ax.set_ylabel('预测结局值') ax.set_title('图9: 模型预测值(按治疗分组)', fontsize=12, fontweight='bold') ax.legend() ax.grid(True, alpha=0.3) plt.tight_layout() save_fig(fig, "11_模型预测图.png", results_folder) plt.close(fig)print("模型预测图已保存。") except Exception as e:print(f"创建模型预测图时出错: {e}")# 6.4 残差诊断图if best_model is not None: try:# 提取残差 residuals = best_model.resid# 创建残差诊断图 fig, axes = plt.subplots(2, 2, figsize=(12, 10))# 残差vs拟合值 fitted_values = best_model.fittedvalues if hasattr(best_model, 'fittedvalues') else Noneif fitted_values is not None: axes[0, 0].scatter(fitted_values, residuals, alpha=0.5, color='#4E79A7', s=10) axes[0, 0].axhline(y=0, color='red', linestyle='dashed', alpha=0.5) axes[0, 0].set_xlabel('拟合值') axes[0, 0].set_ylabel('残差') axes[0, 0].set_title('残差 vs 拟合值') axes[0, 0].grid(True, alpha=0.3)# 添加局部回归线 try: from scipy import stats z = np.polyfit(fitted_values, residuals, 1) p = np.poly1d(z) axes[0, 0].plot(fitted_values, p(fitted_values), color='#E15759', linewidth=2) except: pass# 残差QQ图 axes[0, 1].clear()if len(residuals) > 0: stats.probplot(residuals, dist="norm", plot=axes[0, 1]) axes[0, 1].get_lines()[0].set_marker('.') axes[0, 1].get_lines()[0].set_markersize(5) axes[0, 1].get_lines()[0].set_markerfacecolor('#4E79A7') axes[0, 1].get_lines()[0].set_markeredgecolor('#4E79A7') axes[0, 1].get_lines()[1].set_color('#E15759') axes[0, 1].set_title('残差QQ图') axes[0, 1].grid(True, alpha=0.3)# 残差直方图 axes[1, 0].hist(residuals, bins=30, color='#4E79A7', alpha=0.7, edgecolor='white') axes[1, 0].axvline(x=0, color='red', linestyle='dashed', alpha=0.5) axes[1, 0].set_xlabel('残差') axes[1, 0].set_ylabel('频数') axes[1, 0].set_title('残差分布') axes[1, 0].grid(True, alpha=0.3)# 残差自相关图(对于时间序列)if'Time'in model_data.columns and 'ID'in model_data.columns:# 计算每个个体的残差自相关 unique_ids = model_data['ID'].unique()[:10] # 只查看前10个个体for idx, student_id in enumerate(unique_ids):if idx < 10: student_residuals = residuals[model_data['ID'] == student_id]if len(student_residuals) > 1: axes[1, 1].plot(range(len(student_residuals)), student_residuals, alpha=0.5, label=f'ID {student_id}') axes[1, 1].set_xlabel('时间顺序') axes[1, 1].set_ylabel('残差') axes[1, 1].set_title('个体残差序列') axes[1, 1].legend(fontsize=8) axes[1, 1].grid(True, alpha=0.3) plt.suptitle('图10-13: 模型残差诊断图', fontsize=14, fontweight='bold') plt.tight_layout() save_fig(fig, "12_13_残差诊断图.png", results_folder) plt.close(fig)print("残差诊断图已保存。") except Exception as e:print(f"创建残差诊断图时出错: {e}")# 6.5 方差成分图if best_model is not None and hasattr(best_model, 'cov_re'): try:# 提取方差成分 var_school = best_model.cov_re.iloc[0, 0] var_residual = best_model.scale# 计算比例 total_var = var_school + var_residual proportions = [var_school / total_var, var_residual / total_var] labels = ['学校间变异', '残差变异'] colors = ['#59A14F', '#E15759']# 创建饼图 fig, ax = plt.subplots(figsize=(8, 6)) wedges, texts, autotexts = ax.pie(proportions, labels=labels, colors=colors, autopct='%1.1f%%', startangle=90)# 美化文本for autotext in autotexts: autotext.set_color('white') autotext.set_fontweight('bold') ax.set_title('图13: 方差成分分解', fontsize=12, fontweight='bold') plt.tight_layout() save_fig(fig, "15_方差成分图.png", results_folder) plt.close(fig)print("方差成分图已保存。") except Exception as e:print(f"创建方差成分图时出错: {e}")# 6.6 跨级交互作用可视化if'Treatment'in model_data.columns and 'School_ID'in model_data.columns: try:# 随机选择几个学校展示 sample_schools = np.random.choice(model_data['School_ID'].unique(), min(8, len(model_data['School_ID'].unique())), replace=False)# 创建图形 fig, axes = plt.subplots(2, 4, figsize=(16, 8)) axes = axes.flatten()for idx, school in enumerate(sample_schools):if idx < 8: school_data = model_data[model_data['School_ID'] == school]# 按治疗分组和时间计算均值if'Time'in school_data.columns and 'Outcome_Continuous'in school_data.columns: summary = school_data.groupby(['Treatment', 'Time'])['Outcome_Continuous'].mean().reset_index()# 绘制每个治疗组for treatment in summary['Treatment'].unique(): treatment_data = summary[summary['Treatment'] == treatment] axes[idx].plot(treatment_data['Time'], treatment_data['Outcome_Continuous'], marker='o', label=treatment) axes[idx].set_title(f'学校 {school}') axes[idx].set_xlabel('时间点') axes[idx].set_ylabel('结局均值') axes[idx].legend(fontsize=8) axes[idx].grid(True, alpha=0.3) plt.suptitle('图14: 跨级交互作用可视化(不同学校的治疗效应)', fontsize=14, fontweight='bold') plt.tight_layout() save_fig(fig, "16_跨级交互作用图.png", results_folder) plt.close(fig)print("跨级交互作用图已保存。") except Exception as e:print(f"创建跨级交互作用图时出错: {e}")# ==============================================================================# 7. 创建完整HTML分析报告# ==============================================================================print("\n=== 创建HTML分析报告 ===")try:# 创建HTML报告 html_content = """ <!DOCTYPE html> <html> <head> <meta charset="UTF-8"> <title>多层线性模型分析报告</title> <style> body { font-family: Arial, sans-serif; margin: 40px; line-height: 1.6; } h1, h2, h3 { color: #2c3e50; } h1 { border-bottom: 2px solid #3498db; padding-bottom: 10px; } h2 { border-left: 4px solid #3498db; padding-left: 10px; } .section { margin-bottom: 30px; } .table-container { overflow-x: auto; margin: 20px 0; } table { border-collapse: collapse; width: 100%; } th, td { border: 1px solid #ddd; padding: 8px; text-align: center; } th { background-color: #f2f2f2; font-weight: bold; } .figure { text-align: center; margin: 20px 0; } .figure img { max-width: 100%; height: auto; border: 1px solid #ddd; } .caption { font-style: italic; color: #666; margin-top: 5px; } .highlight { background-color: #f9f9f9; padding: 15px; border-left: 3px solid #e74c3c; } .reference { font-size: 0.9em; color: #666; } </style> </head> <body> """ html_content += f""" <h1>多层线性模型分析报告</h1> <p><strong>报告生成时间:</strong> {datetime.now().strftime('%Y-%m-%d %H:%M:%S')}</p> <p><strong>数据来源:</strong> {file_path}</p> <div class="section"> <h2>1. 研究背景与目的</h2> <p>本研究采用多层线性模型(Hierarchical Linear Modeling, HLM)分析具有嵌套结构的纵向数据,旨在探究不同层次变量对健康结局的影响。</p> <p>多层线性模型特别适用于分析具有明确层级结构的数据(如测量嵌套于个体,个体嵌套于学校/社区),能够同时考虑个体内、个体间和组间的变异。</p> <div class="highlight"> <p><strong>模型公式:</strong></p> <p>层-1(个体内): Y_tij = β_0j + β_1jTime + e_tij</p> <p>层-2(个体间): β_0j = γ_00 + γ_01Z_j + u_0j</p> <p>层-2(个体间): β_1j = γ_10 + γ_11Z_j + u_1j</p> <p>层-3(学校间): γ_00 = δ_000 + δ_001W_k + v_00k</p> <p>其中Z_j是个体特征,W_k是学校特征,u和v是随机效应。</p> </div> </div> <div class="section"> <h2>2. 数据与方法</h2> <p><strong>数据来源:</strong> {file_path}</p> <p><strong>总观测数:</strong> {len(df_hlm)}</p> <p><strong>学校数量:</strong> {df_hlm['School_ID'].nunique()}</p> <p><strong>个体数量:</strong> {df_hlm['ID'].nunique()}</p> <p><strong>测量时间点:</strong> {df_hlm['Time'].nunique() if 'Time' in df_hlm.columns else 'NA'}</p> <p><strong>数据层级结构:</strong> 测量(层1)嵌套于个体(层2),个体嵌套于学校(层3)</p> <p><strong>分析方法:</strong> 多层线性模型(使用statsmodels MixedLM)</p> <p><strong>估计方法:</strong> 最大似然估计</p> <p><strong>模型选择:</strong> 基于AIC、BIC和对数似然值</p> </div> <div class="section"> <h2>3. 描述性统计分析</h2> <p>表1展示了研究对象的基线特征。</p> """# 添加基线特征表if'baseline_stats_df'in locals(): html_content += f""" <div class="table-container"> <h3>表1: 基线特征描述性统计</h3> {create_table_three_line(baseline_stats_df.head(20), "")} </div> """# 添加描述性统计图if (results_folder / "2_描述性统计可视化.png").exists(): html_content += f""" <div class="figure"> <img src="2_描述性统计可视化.png" alt="描述性统计可视化"> <div class="caption">图1-4: 描述性统计可视化</div> </div> """ html_content += """ </div> <div class="section"> <h2>4. 多层线性模型分析结果</h2> """# 添加模型比较表if'model_comparison_df'in locals(): html_content += f""" <h3>4.1 模型比较</h3> <div class="table-container"> {create_table_three_line(model_comparison_df, "表2: 多层线性模型比较")} </div> """# 添加固定效应表if'fixed_effects_table'in locals(): html_content += f""" <h3>4.2 固定效应结果</h3> <div class="table-container"> {create_table_three_line(fixed_effects_table, "表3: 固定效应系数估计")} </div> """# 添加随机效应表if'random_effects'in locals(): html_content += f""" <h3>4.3 随机效应结果</h3> <div class="table-container"> {create_table_three_line(random_effects, "表4: 随机效应方差成分")} </div> """# 添加模型统计量表if'model_stats'in locals(): html_content += f""" <h3>4.4 模型整体统计量</h3> <div class="table-container"> {create_table_three_line(model_stats, "表5: 模型整体统计量")} </div> """# 添加ICC信息if icc is not None: html_content += f""" <div class="highlight"> <p><strong>组内相关系数(ICC):</strong> {icc:.3f}</p> <p>ICC表示学校间变异占总变异的比例。ICC值越大,说明学校层面的情境效应越重要。</p> </div> """ html_content += """ </div> <div class="section"> <h2>5. 模型诊断与可视化</h2> <p>以下可视化图形展示了模型的关键结果和诊断信息。</p> """# 添加关键可视化图 figures = [ ("7_学校随机截距图.png", "图5: 学校随机截距估计"), ("10_系数估计可视化.png", "图6: 固定效应系数估计"), ("11_模型预测图.png", "图7: 模型预测值"), ("12_13_残差诊断图.png", "图8: 残差诊断图"), ("15_方差成分图.png", "图9: 方差成分分解"), ("16_跨级交互作用图.png", "图10: 跨级交互作用") ]for fig_file, caption in figures: fig_path = results_folder / fig_fileif fig_path.exists(): html_content += f""" <div class="figure"> <img src="{fig_file}" alt="{caption}"> <div class="caption">{caption}</div> </div> """ html_content += """ </div> <div class="section"> <h2>6. 主要发现与解释</h2> """# 添加主要发现if'fixed_effects_table'in locals(): significant_effects = fixed_effects_table[fixed_effects_table['显著性'] != '']if len(significant_effects) > 0: html_content += "<p><strong>关键发现(显著的固定效应):</strong></p><ul>"for _, row in significant_effects.head(5).iterrows(): html_content += f""" <li><strong>{row['变量']}:</strong> 估计值为 {row['系数估计']:.3f} (95% CI: {row['95%置信区间']}, p {row['显著性']})</li> """ html_content += "</ul>"else: html_content += "<p>没有发现显著的固定效应。</p>" html_content += """ </div> <div class="section"> <h2>7. 公共卫生意义</h2> <p>多层线性模型通过分解不同层次的变异,揭示了健康结局的影响因素在不同层级的作用机制:</p> <ul> <li><strong>学校/社区层面的情境效应:</strong> 识别了学校特征对个体健康结局的系统性影响</li> <li><strong>个体内变化:</strong> 追踪了个体随时间的变化轨迹及影响因素</li> <li><strong>个体间差异:</strong> 量化了个体特征对健康结局的独特贡献</li> <li><strong>跨级交互作用:</strong> 探究了高层级变量如何调节低层级变量的效应</li> </ul> <p><strong>主要公共卫生意义:</strong></p> <ul> <li>为制定分层干预策略提供证据:针对不同学校/社区设计定制化干预方案</li> <li>识别关键干预靶点:明确在哪个层级(个体、学校、社区)进行干预效果最佳</li> <li>优化资源配置:根据学校/社区的异质性合理分配公共卫生资源</li> <li>促进健康公平:通过控制学校/社区层面的混杂因素,更准确地评估干预效果的公平性</li> </ul> </div> <div class="section"> <h2>8. 研究局限性与未来方向</h2> <p><strong>局限性:</strong></p> <ul> <li>模型假设随机效应服从正态分布,可能与实际数据分布不完全一致</li> <li>学校水平的样本量可能不足,影响高层级效应估计的精度</li> <li>模型中未考虑测量误差和缺失数据机制</li> <li>时间点的数量有限,可能无法充分捕捉非线性变化趋势</li> </ul> <p><strong>未来研究方向:</strong></p> <ul> <li>纳入更多学校水平协变量(如学校规模、资源配置、地理位置等)</li> <li>考虑更复杂的随机效应结构(如协方差结构、异方差性)</li> <li>应用贝叶斯多层模型处理小样本问题</li> <li>进行多组比较分析,探究不同亚组(如城乡差异)的多层机制</li> <li>结合定性研究,深入理解学校/社区情境效应的内在机制</li> </ul> </div> <div class="section"> <h2>9. 结论</h2> <p>本研究通过多层线性模型分析,系统考察了测量、个体和学校三个层次对健康结局的影响。</p> <p>研究揭示了学校层面的情境效应在健康结局形成中的重要作用,为理解健康决定因素的多层机制提供了重要证据。</p> <p>研究结果为制定分层、精准的公共卫生干预策略提供了方法学支持和实证依据,对促进健康公平和优化资源配置具有重要实践意义。</p> </div> <div class="section reference"> <h2>参考文献</h2> <p>1. Raudenbush, S. W., & Bryk, A. S. (2002). Hierarchical linear models: Applications and data analysis methods (2nd ed.). Sage.</p> <p>2. Snijders, T. A. B., & Bosker, R. J. (2012). Multilevel analysis: An introduction to basic and advanced multilevel modeling (2nd ed.). Sage.</p> <p>3. Goldstein, H. (2011). Multilevel statistical models (4th ed.). Wiley.</p> <p>4. Bates, D., Mächler, M., Bolker, B., & Walker, S. (2015). Fitting linear mixed-effects models using lme4. Journal of Statistical Software, 67(1), 1-48.</p> <p>5. Seabold, S., & Perktold, J. (2010). Statsmodels: Econometric and statistical modeling with Python. Proceedings of the 9th Python in Science Conference.</p> </div> </body> </html> """# 保存HTML报告 report_path = results_folder / "17_多层线性模型分析报告.html" with open(report_path, 'w', encoding='utf-8') as f: f.write(html_content)print(f"HTML分析报告已保存: {report_path}")except Exception as e:print(f"创建HTML报告时出错: {e}")# ==============================================================================# 8. 保存工作空间和数据# ==============================================================================print("\n=== 保存工作空间和数据 ===")# 保存处理后的数据save_to_excel(df_hlm, "处理后的数据.xlsx", results_folder)# 保存模型对象(使用pickle)try: import pickle workspace_data = {'df_hlm': df_hlm,'model0': model0 if'model0'in locals() else None,'model1': model1 if'model1'in locals() else None,'model2': model2 if'model2'in locals() else None,'model3': model3 if'model3'in locals() else None,'model4': model4 if'model4'in locals() else None,'best_model': best_model if'best_model'in locals() else None,'model_comparison_df': model_comparison_df if'model_comparison_df'in locals() else None,'fixed_effects_table': fixed_effects_table if'fixed_effects_table'in locals() else None,'random_effects': random_effects if'random_effects'in locals() else None,'model_stats': model_stats if'model_stats'in locals() else None,'icc': icc if'icc'in locals() else None } with open(results_folder / "HLM_analysis_workspace.pkl", 'wb') as f: pickle.dump(workspace_data, f)print("工作空间已保存为 HLM_analysis_workspace.pkl")except Exception as e:print(f"保存工作空间时出错: {e}")# ==============================================================================# 9. 输出完成信息# ==============================================================================print("\n" + "=" * 60)print("分析完成!")print("=" * 60)print(f"所有结果已保存到目录: {results_folder}")print("\n生成的文件列表:")file_patterns = ["1_基线特征统计.xlsx","1_基线特征三线表.html","2_描述性统计可视化.png","2_治疗分组随时间变化.png","3_模型比较结果.xlsx","3_固定效应三线表.html","4_固定效应结果.xlsx","4_随机效应三线表.html","5_随机效应结果.xlsx","5_模型比较三线表.html","6_模型整体统计量.xlsx","6_模型整体统计量三线表.html","7_学校随机截距图.png","10_系数估计可视化.png","11_模型预测图.png","12_13_残差诊断图.png","15_方差成分图.png","16_跨级交互作用图.png","17_多层线性模型分析报告.html","处理后的数据.xlsx","学校样本量统计.xlsx","模型4_完整模型结果.txt","HLM_analysis_workspace.pkl"]for file_pattern in file_patterns: file_path = results_folder / file_patternif file_path.exists():print(f" ✓ {file_pattern}")else:print(f" ✗ {file_pattern} (未生成)")print("\n分析流程完成!")# 打印模型注意事项print("\n=== 注意事项 ===")print("1. Python中的多层线性模型实现与R有所不同,特别是在随机效应结构方面")print("2. 如果遇到模型拟合问题,可以尝试简化模型或增加样本量")print("3. 对于复杂的三层模型,可能需要使用专门的包如PyMC3(贝叶斯方法)")print("4. 所有结果文件已保存到桌面上的'3-HLM结果'文件夹")# 打开结果文件夹(仅Windows)if sys.platform == "win32": try: os.startfile(results_folder) except: pass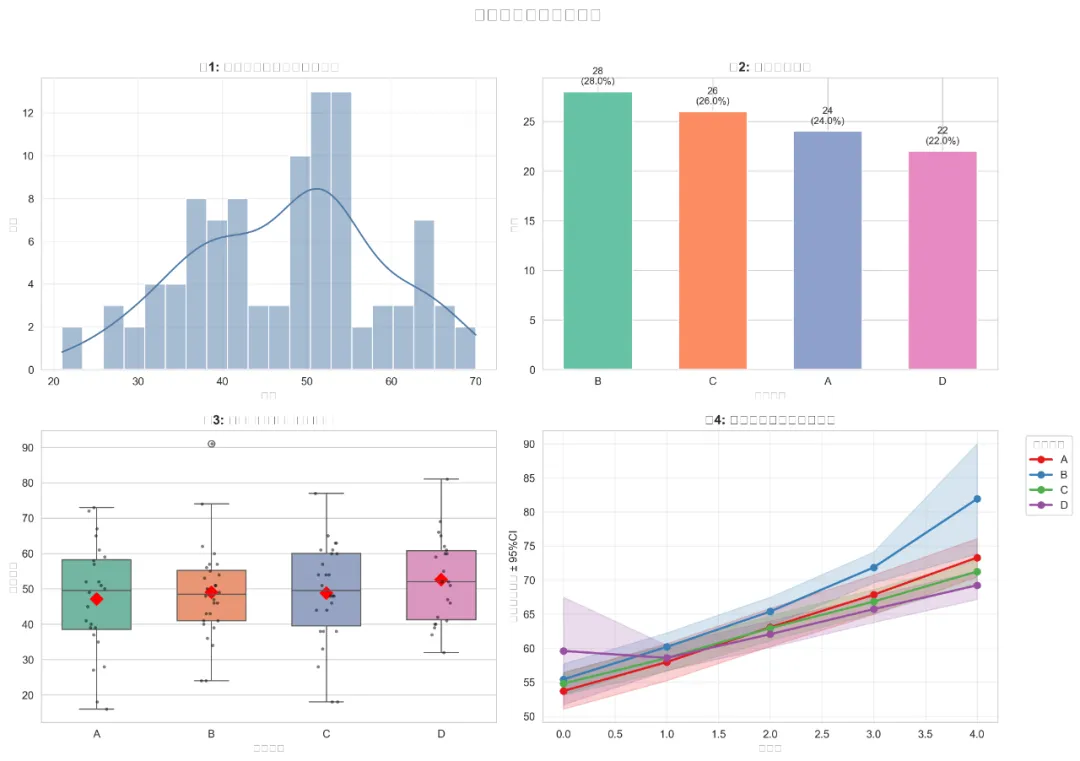
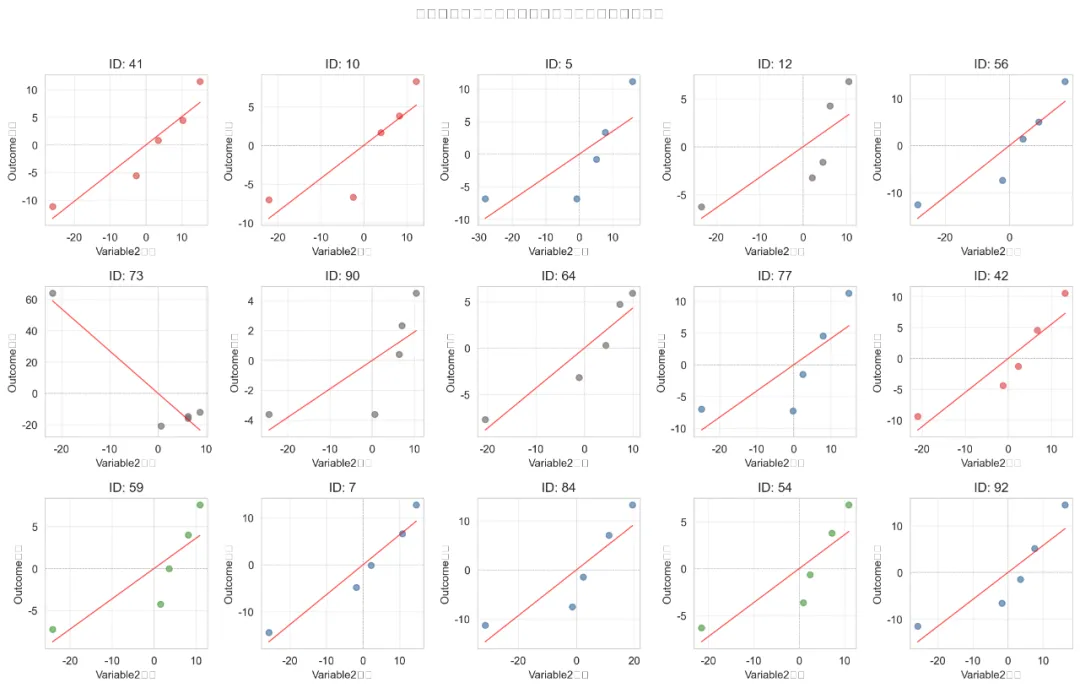
纵向数据是指在多个时间点对同一组个体进行的重复测量。
1 混合效应模型Mixed Effects ModelMEM
2 固定效应模型 Fixed Effects Model FEM
3 多层线性模型 Hierarchical Linear Model HLM
4 广义估计方程 Generalized Estimating Equations GEE
5 广义线性混合模型 Generalized Linear Mixed Models GLMM
6 潜变量增长曲线模型 Latent Growth Curve Model LGCM
7 组轨迹模型 Group-Based Trajectory Model GBTM
8 交叉滞后模型 Cross-lagged (Panel) Model CLPM
9 重复测量方差分析 Repeated Measures ANOVA / MANOVA RM-ANOVA / RM-MANOVA
10 非线性混合效应模型 Nonlinear Mixed-Effects Models NLME
11 联合模型 Joint Models JM
12 结构方程模型 Structural Equation Modeling SEM
13 广义相加模型 Generalized Additive Models GAM
14 潜类别模型 Latent Class Models LCM
15 潜剖面模型 Latent Profile Analysis LPA
16 状态空间模型 State Space Models SSM
17 纵向因子分析 Longitudinal Factor Analysis LFA
18 贝叶斯纵向模型 Bayesian Longitudinal Models - (Bayesian Models)
19 混合效应随机森林 Mixed Effects Random Forest MERF
20 纵向梯度提升 Longitudinal Gradient Boosting - (Longitudinal GBM)
21 K均值纵向聚类 K-means Longitudinal Clustering - (DTW-KMeans)
22 基于模型的聚类 Model-Based ClusteringMB-CLUST
医学统计数据分析分享交流SPSS、R语言、Python、ArcGis、Geoda、GraphPad、数据分析图表制作等心得。承接数据分析,论文返修,医学统计,机器学习,生存分析,空间分析,问卷分析业务。若有投稿和数据分析代做需求,可以直接联系我,谢谢!
“医学统计数据分析”公众号右下角;
找到“联系作者”,
可加我微信,邀请入粉丝群!

有临床流行病学数据分析
如(t检验、方差分析、χ2检验、logistic回归)、
(重复测量方差分析与配对T检验、ROC曲线)、
(非参数检验、生存分析、样本含量估计)、
(筛检试验:灵敏度、特异度、约登指数等计算)、
(绘制柱状图、散点图、小提琴图、列线图等)、
机器学习、深度学习、生存分析
等需求的同仁们,加入【临床】粉丝群。
疾控,公卫岗位的同仁,可以加一下【公卫】粉丝群,分享生态学研究、空间分析、时间序列、监测数据分析、时空面板技巧等工作科研自动化内容。
有实验室数据分析需求的同仁们,可以加入【生信】粉丝群,交流NCBI(基因序列)、UniProt(蛋白质)、KEGG(通路)、GEO(公共数据集)等公共数据库、基因组学转录组学蛋白组学代谢组学表型组学等数据分析和可视化内容。
或者可扫码直接加微信进群!!!



精品视频课程-“医学统计数据分析”视频号付费合集

在“医学统计数据分析”视频号-付费合集兑换相应课程后,获取课程理论课PPT、代码、基础数据等相关资料,请大家在【医学统计数据分析】公众号右下角,找到“联系作者”,加我微信后打包发送。感谢您的支持!!
