# pip install tensorflow pandas numpy matplotlib seaborn scikit-learnimport osimport pandas as pdimport numpy as npimport matplotlib.pyplot as pltimport seaborn as snsfrom datetime import datetime, timedeltaimport warningswarnings.filterwarnings('ignore')# 深度学习相关库import tensorflow as tffrom tensorflow import kerasfrom tensorflow.keras.models import Sequentialfrom tensorflow.keras.layers import LSTM, Dense, Dropoutfrom tensorflow.keras.optimizers import Adamfrom tensorflow.keras.callbacks import EarlyStopping# 数据预处理和可视化from sklearn.preprocessing import StandardScalerfrom sklearn.metrics import mean_absolute_error, mean_squared_error, r2_scoreimport scipy.stats as stats# 设置中文字体plt.rcParams['font.sans-serif'] = ['SimHei']plt.rcParams['axes.unicode_minus'] = Falsedef get_desktop_path():"""获取桌面路径"""return os.path.join(os.path.expanduser("~"), "Desktop")def create_directory(path):"""创建目录"""if not os.path.exists(path): os.makedirs(path)return pathclass DiseaseRNNModel:"""疾病RNN预测模型""" def __init__(self, results_path): self.results_path = results_path self.scalers = {} self.models = {} self.results = {} def prepare_data(self, data, target_var, feature_vars, lookback=60, forecast_horizon=1, train_start='1981-01-01', train_end='2015-12-31', test_start='2016-01-01', test_end='2025-12-31'):"""准备RNN数据"""# 筛选数据 df = data[['timestamp', target_var] + feature_vars].copy() df['timestamp'] = pd.to_datetime(df['timestamp']) df = df.sort_values('timestamp').reset_index(drop=True)# 划分训练集和测试集 train_data = df[(df['timestamp'] >= train_start) & (df['timestamp'] <= train_end)] test_data = df[(df['timestamp'] >= test_start) & (df['timestamp'] <= test_end)]# 标准化数据 scaler_target = StandardScaler() scaler_features = StandardScaler()# 训练集标准化 train_scaled = train_data.copy() train_scaled[target_var] = scaler_target.fit_transform(train_data[[target_var]]) train_scaled[feature_vars] = scaler_features.fit_transform(train_data[feature_vars])# 测试集标准化 test_scaled = test_data.copy() test_scaled[target_var] = scaler_target.transform(test_data[[target_var]]) test_scaled[feature_vars] = scaler_features.transform(test_data[feature_vars])# 创建时间序列样本 def create_sequences(data, lookback, forecast_horizon): X, y = [], [] dates = []for i in range(lookback, len(data) - forecast_horizon + 1): X.append(data.iloc[i - lookback:i][[target_var] + feature_vars].values) y.append(data.iloc[i + forecast_horizon - 1][target_var]) dates.append(data.iloc[i]['timestamp'])return np.array(X), np.array(y), dates# 创建序列 X_train, y_train, train_dates = create_sequences(train_scaled, lookback, forecast_horizon) X_test, y_test, test_dates = create_sequences(test_scaled, lookback, forecast_horizon)return {'X_train': X_train, 'y_train': y_train, 'train_dates': train_dates,'X_test': X_test, 'y_test': y_test, 'test_dates': test_dates,'scaler_target': scaler_target, 'scaler_features': scaler_features,'original_train': train_data.iloc[lookback:len(train_data) - forecast_horizon + 1],'original_test': test_data.iloc[lookback:len(test_data) - forecast_horizon + 1] } def build_model(self, input_shape, units=50, dropout_rate=0.2):"""构建LSTM模型""" model = Sequential([ LSTM(units, return_sequences=True, input_shape=input_shape), Dropout(dropout_rate), LSTM(units, return_sequences=False), Dropout(dropout_rate), Dense(25, activation='relu'), Dense(1, activation='linear') ]) model.compile( optimizer=Adam(learning_rate=0.001), loss='mse', metrics=['mae'] )return model def calculate_metrics(self, y_true, y_pred):"""计算评估指标""" mae = mean_absolute_error(y_true, y_pred) mse = mean_squared_error(y_true, y_pred) rmse = np.sqrt(mse) mape = np.mean(np.abs((y_true - y_pred) / y_true)) * 100 r2 = r2_score(y_true, y_pred)return {'MAE': mae, 'MSE': mse, 'RMSE': rmse,'MAPE': mape, 'R2': r2 } def train_model(self, disease, data, feature_vars, lookback=60, epochs=150, batch_size=64):"""训练单个疾病模型"""print(f"正在处理疾病: {disease}")# 准备数据 rnn_data = self.prepare_data(data, disease, feature_vars, lookback=lookback)# 构建模型 model = self.build_model( input_shape=(lookback, len([disease] + feature_vars)) )print("模型结构:") model.summary()# 设置早停法 early_stop = EarlyStopping( monitor='val_loss', patience=20, restore_best_weights=True )# 训练模型history = model.fit( rnn_data['X_train'], rnn_data['y_train'], epochs=epochs, batch_size=batch_size, validation_split=0.2, verbose=1, callbacks=[early_stop] )# 预测 train_predictions = model.predict(rnn_data['X_train']) test_predictions = model.predict(rnn_data['X_test'])# 反标准化 train_pred_unscaled = rnn_data['scaler_target'].inverse_transform(train_predictions).flatten() test_pred_unscaled = rnn_data['scaler_target'].inverse_transform(test_predictions).flatten() train_true_unscaled = rnn_data['original_train'][disease].values test_true_unscaled = rnn_data['original_test'][disease].values# 计算评估指标 train_metrics = self.calculate_metrics(train_true_unscaled, train_pred_unscaled) test_metrics = self.calculate_metrics(test_true_unscaled, test_pred_unscaled)# 保存结果 self.results[disease] = {'disease': disease,'model': model,'history': history,'train_dates': rnn_data['train_dates'],'test_dates': rnn_data['test_dates'],'train_true': train_true_unscaled,'train_pred': train_pred_unscaled,'test_true': test_true_unscaled,'test_pred': test_pred_unscaled,'train_metrics': train_metrics,'test_metrics': test_metrics,'lookback': lookback }return self.results[disease]def plot_training_history(results, target_diseases, save_dir):"""绘制训练历史""" create_directory(save_dir)for disease in target_diseases: result = results[disease]history = result['history'] fig, ax = plt.subplots(figsize=(10, 6)) ax.plot(history.history['loss'], label='训练损失', linewidth=2) ax.plot(history.history['val_loss'], label='验证损失', linewidth=2) ax.set_title(f'{disease} - RNN训练历史', fontsize=14, fontweight='bold') ax.set_xlabel('训练轮次') ax.set_ylabel('损失值') ax.legend() ax.grid(True, alpha=0.3) plt.tight_layout() plt.savefig(f'{save_dir}/Training_History_{disease}.png', dpi=300, bbox_inches='tight') plt.close()
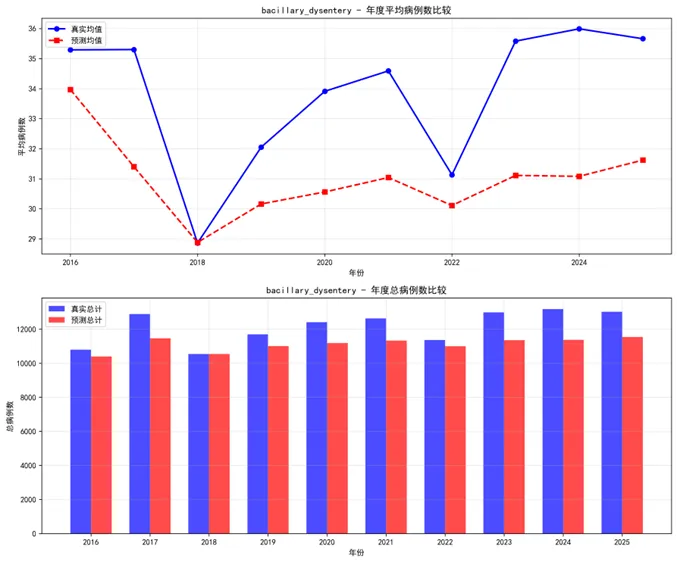
def plot_prediction_comparison(results, target_diseases, save_dir):"""绘制预测结果对比""" create_directory(save_dir)for disease in target_diseases: result = results[disease]# 训练集预测对比 fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(14, 10))# 训练集 ax1.plot(result['train_dates'], result['train_true'], label='真实值', linewidth=1.5, color='blue') ax1.plot(result['train_dates'], result['train_pred'], label='预测值', linewidth=1.5, linestyle='--', color='red') ax1.set_title(f'{disease} - 训练集预测对比 (1981-2015)', fontsize=12, fontweight='bold') ax1.set_ylabel('病例数') ax1.legend() ax1.grid(True, alpha=0.3)# 测试集 ax2.plot(result['test_dates'], result['test_true'], label='真实值', linewidth=1.5, color='blue') ax2.plot(result['test_dates'], result['test_pred'], label='预测值', linewidth=1.5, linestyle='--', color='red') ax2.set_title(f'{disease} - 测试集预测对比 (2016-2025)', fontsize=12, fontweight='bold') ax2.set_xlabel('日期') ax2.set_ylabel('病例数') ax2.legend() ax2.grid(True, alpha=0.3) plt.tight_layout() plt.savefig(f'{save_dir}/Prediction_Comparison_{disease}.png', dpi=300, bbox_inches='tight') plt.close()
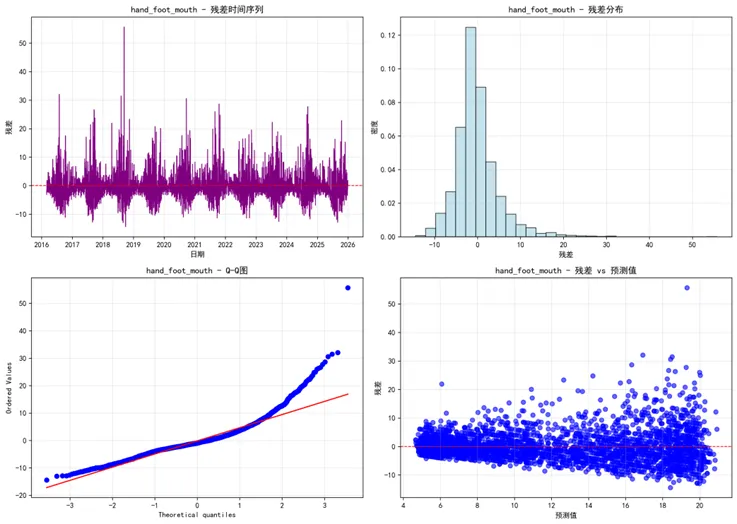
def plot_residual_analysis(results, target_diseases, save_dir):"""绘制残差分析""" create_directory(save_dir)for disease in target_diseases: result = results[disease] residuals = result['test_true'] - result['test_pred'] fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, figsize=(14, 10))# 残差时间序列 ax1.plot(result['test_dates'], residuals, color='purple', linewidth=1) ax1.axhline(y=0, color='red', linestyle='--', linewidth=1) ax1.set_title(f'{disease} - 残差时间序列') ax1.set_xlabel('日期') ax1.set_ylabel('残差') ax1.grid(True, alpha=0.3)# 残差分布 ax2.hist(residuals, bins=30, density=True, alpha=0.7, color='lightblue', edgecolor='black') ax2.set_title(f'{disease} - 残差分布') ax2.set_xlabel('残差') ax2.set_ylabel('密度') ax2.grid(True, alpha=0.3)# Q-Q图 stats.probplot(residuals, dist="norm", plot=ax3) ax3.set_title(f'{disease} - Q-Q图') ax3.grid(True, alpha=0.3)# 残差vs预测值 ax4.scatter(result['test_pred'], residuals, alpha=0.6, color='blue') ax4.axhline(y=0, color='red', linestyle='--', linewidth=1) ax4.set_title(f'{disease} - 残差 vs 预测值') ax4.set_xlabel('预测值') ax4.set_ylabel('残差') ax4.grid(True, alpha=0.3) plt.tight_layout() plt.savefig(f'{save_dir}/Residual_Analysis_{disease}.png', dpi=300, bbox_inches='tight') plt.close()def plot_performance_comparison(results, target_diseases, save_dir):"""绘制性能比较图""" create_directory(save_dir)# 收集性能数据 performance_data = []for disease in target_diseases: result = results[disease]# 训练集指标 performance_data.append({'Disease': disease, 'Dataset': '训练集','MAE': result['train_metrics']['MAE'],'RMSE': result['train_metrics']['RMSE'],'MAPE': result['train_metrics']['MAPE'],'R2': result['train_metrics']['R2'] })# 测试集指标 performance_data.append({'Disease': disease, 'Dataset': '测试集','MAE': result['test_metrics']['MAE'],'RMSE': result['test_metrics']['RMSE'],'MAPE': result['test_metrics']['MAPE'],'R2': result['test_metrics']['R2'] }) perf_df = pd.DataFrame(performance_data)# 绘制性能比较图 fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, figsize=(16, 12))# MAE比较 sns.barplot(data=perf_df, x='Disease', y='MAE', hue='Dataset', ax=ax1, alpha=0.8) ax1.set_title('RNN模型MAE比较') ax1.tick_params(axis='x', rotation=45)# RMSE比较 sns.barplot(data=perf_df, x='Disease', y='RMSE', hue='Dataset', ax=ax2, alpha=0.8) ax2.set_title('RNN模型RMSE比较') ax2.tick_params(axis='x', rotation=45)# MAPE比较 sns.barplot(data=perf_df, x='Disease', y='MAPE', hue='Dataset', ax=ax3, alpha=0.8) ax3.set_title('RNN模型MAPE比较') ax3.tick_params(axis='x', rotation=45)# R²比较 sns.barplot(data=perf_df, x='Disease', y='R2', hue='Dataset', ax=ax4, alpha=0.8) ax4.set_title('RNN模型R²比较') ax4.set_ylim(0, 1) ax4.tick_params(axis='x', rotation=45) plt.tight_layout() plt.savefig(f'{save_dir}/Performance_Comparison_All.png', dpi=300, bbox_inches='tight') plt.close()return perf_dfdef plot_prediction_intervals(results, target_diseases, save_dir):"""绘制预测区间""" create_directory(save_dir)for disease in target_diseases: result = results[disease] residuals = result['test_true'] - result['test_pred'] residual_sd = np.std(residuals) prediction_data = pd.DataFrame({'Date': result['test_dates'],'True': result['test_true'],'Predicted': result['test_pred'],'Lower_95': result['test_pred'] - 1.96 * residual_sd,'Upper_95': result['test_pred'] + 1.96 * residual_sd,'Lower_80': result['test_pred'] - 1.28 * residual_sd,'Upper_80': result['test_pred'] + 1.28 * residual_sd }) fig, ax = plt.subplots(figsize=(14, 8))# 绘制预测区间 ax.fill_between(prediction_data['Date'], prediction_data['Lower_95'], prediction_data['Upper_95'], alpha=0.3, label='95% 预测区间', color='lightblue') ax.fill_between(prediction_data['Date'], prediction_data['Lower_80'], prediction_data['Upper_80'], alpha=0.5, label='80% 预测区间', color='lightgreen')# 绘制真实值和预测值 ax.plot(prediction_data['Date'], prediction_data['True'], label='真实值', linewidth=1.5, color='blue') ax.plot(prediction_data['Date'], prediction_data['Predicted'], label='预测值', linewidth=1.5, color='red') ax.set_title(f'{disease} - 预测区间可视化', fontsize=14, fontweight='bold') ax.set_xlabel('日期') ax.set_ylabel('病例数') ax.legend() ax.grid(True, alpha=0.3) plt.tight_layout() plt.savefig(f'{save_dir}/Prediction_Interval_{disease}.png', dpi=300, bbox_inches='tight') plt.close()def plot_annual_comparison(results, target_diseases, save_dir):"""绘制年度比较图""" create_directory(save_dir)for disease in target_diseases: result = results[disease] annual_data = pd.DataFrame({'Date': result['test_dates'],'True': result['test_true'],'Predicted': result['test_pred'] }) annual_data['Year'] = pd.to_datetime(annual_data['Date']).dt.year annual_summary = annual_data.groupby('Year').agg({'True': ['mean', 'sum'],'Predicted': ['mean', 'sum'] }).round(2) annual_summary.columns = ['True_Mean', 'True_Total', 'Predicted_Mean', 'Predicted_Total'] annual_summary = annual_summary.reset_index() fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(12, 10))# 年度均值比较 ax1.plot(annual_summary['Year'], annual_summary['True_Mean'], label='真实均值', marker='o', linewidth=2, color='blue') ax1.plot(annual_summary['Year'], annual_summary['Predicted_Mean'], label='预测均值', marker='s', linestyle='--', linewidth=2, color='red') ax1.set_title(f'{disease} - 年度平均病例数比较') ax1.set_xlabel('年份') ax1.set_ylabel('平均病例数') ax1.legend() ax1.grid(True, alpha=0.3)# 年度总计比较 x = np.arange(len(annual_summary['Year'])) width = 0.35 ax2.bar(x - width / 2, annual_summary['True_Total'], width, label='真实总计', alpha=0.7, color='blue') ax2.bar(x + width / 2, annual_summary['Predicted_Total'], width, label='预测总计', alpha=0.7, color='red') ax2.set_title(f'{disease} - 年度总病例数比较') ax2.set_xlabel('年份') ax2.set_ylabel('总病例数') ax2.set_xticks(x) ax2.set_xticklabels(annual_summary['Year']) ax2.legend() ax2.grid(True, alpha=0.3) plt.tight_layout() plt.savefig(f'{save_dir}/Annual_Comparison_{disease}.png', dpi=300, bbox_inches='tight') plt.close()def plot_seasonal_patterns(results, target_diseases, save_dir):"""绘制季节性模式""" create_directory(save_dir)for disease in target_diseases: result = results[disease] seasonal_data = pd.DataFrame({'Date': result['test_dates'],'True': result['test_true'],'Predicted': result['test_pred'] }) seasonal_data['Month'] = pd.to_datetime(seasonal_data['Date']).dt.month monthly_pattern = seasonal_data.groupby('Month').agg({'True': 'mean','Predicted': 'mean' }).reset_index() fig, ax = plt.subplots(figsize=(10, 6)) ax.plot(monthly_pattern['Month'], monthly_pattern['True'], label='真实值', marker='o', linewidth=2, color='blue') ax.plot(monthly_pattern['Month'], monthly_pattern['Predicted'], label='预测值', marker='s', linestyle='--', linewidth=2, color='red') ax.set_title(f'{disease} - 月度季节性模式比较') ax.set_xlabel('月份') ax.set_ylabel('平均病例数') ax.set_xticks(range(1, 13)) ax.legend() ax.grid(True, alpha=0.3) plt.tight_layout() plt.savefig(f'{save_dir}/Seasonal_Pattern_{disease}.png', dpi=300, bbox_inches='tight') plt.close()def save_models_and_results(results, target_diseases, save_dir):"""保存模型和结果""" create_directory(save_dir)# 保存性能汇总 performance_summary = []for disease in target_diseases: result = results[disease] summary = {'Disease': disease,'Train_MAE': result['train_metrics']['MAE'],'Train_RMSE': result['train_metrics']['RMSE'],'Train_MAPE': result['train_metrics']['MAPE'],'Train_R2': result['train_metrics']['R2'],'Test_MAE': result['test_metrics']['MAE'],'Test_RMSE': result['test_metrics']['RMSE'],'Test_MAPE': result['test_metrics']['MAPE'],'Test_R2': result['test_metrics']['R2'],'Lookback_Days': result['lookback'] } performance_summary.append(summary) perf_df = pd.DataFrame(performance_summary) perf_df.to_csv(f'{save_dir}/RNN_Performance_Summary.csv', index=False)# 保存模型for disease in target_diseases: result = results[disease] result['model'].save(f'{save_dir}/RNN_Model_{disease}.h5')return perf_dfdef generate_reports(target_diseases, save_dir):"""生成分析报告""" create_directory(save_dir) analysis_report = pd.DataFrame({'分析项目': ['训练历史', '预测对比', '残差分析', '性能比较','预测区间', '年度比较', '季节性模式', '模型保存'],'图表数量': [len(target_diseases), len(target_diseases), len(target_diseases), 1, len(target_diseases), len(target_diseases), len(target_diseases), 1],'文件位置': ['Training_History', 'Prediction_Comparison', 'Residual_Analysis', 'Performance_Comparison','Prediction_Intervals', 'Annual_Comparison', 'Seasonal_Patterns', 'Saved_Models'],'描述': ['RNN训练损失变化过程', '真实值与预测值对比', '残差分布和模式分析', '各疾病模型性能指标比较','预测不确定性区间展示', '年度汇总统计比较', '季节性模式捕捉能力', '训练好的模型文件'] }) analysis_report.to_csv(f'{save_dir}/RNN_Analysis_Report.csv', index=False, encoding='utf-8-sig')return analysis_reportdef main():"""主函数"""print("开始RNN建模分析...")# 获取桌面路径并创建结果文件夹 desktop_path = get_desktop_path() results_path = create_directory(os.path.join(desktop_path, "Results时间RNN")) data_path = os.path.join(desktop_path, "Results", "combined_weather_disease_data.csv")# 检查数据文件是否存在if not os.path.exists(data_path):print(f"错误: 数据文件不存在: {data_path}")return# 读取数据print("读取数据...") combined_data = pd.read_csv(data_path) combined_data['timestamp'] = pd.to_datetime(combined_data['timestamp'])# 选择要分析的疾病和气象特征 target_diseases = ["influenza", "common_cold", "pneumonia","bacillary_dysentery", "hand_foot_mouth"] weather_features = ["temp_mean", "humidity_mean", "pressure_mean","precipitation_total", "sunshine_hours"]# 初始化模型 rnn_model = DiseaseRNNModel(results_path)# 训练模型print("开始训练RNN模型...")for disease in target_diseases: rnn_model.train_model( disease=disease, data=combined_data, feature_vars=weather_features, lookback=60, epochs=150, batch_size=64 ) results = rnn_model.results# 生成各种可视化print("生成可视化图表...")# 1. 训练历史 plot_training_history(results, target_diseases, os.path.join(results_path, "Training_History"))# 2. 预测对比 plot_prediction_comparison(results, target_diseases, os.path.join(results_path, "Prediction_Comparison"))# 3. 残差分析 plot_residual_analysis(results, target_diseases, os.path.join(results_path, "Residual_Analysis"))# 4. 性能比较 perf_df = plot_performance_comparison(results, target_diseases, os.path.join(results_path, "Performance_Comparison"))# 5. 预测区间 plot_prediction_intervals(results, target_diseases, os.path.join(results_path, "Prediction_Intervals"))# 6. 年度比较 plot_annual_comparison(results, target_diseases, os.path.join(results_path, "Annual_Comparison"))# 7. 季节性模式 plot_seasonal_patterns(results, target_diseases, os.path.join(results_path, "Seasonal_Patterns"))# 8. 保存模型和结果 performance_summary = save_models_and_results(results, target_diseases, os.path.join(results_path, "Saved_Models"))# 9. 生成报告 analysis_report = generate_reports(target_diseases, os.path.join(results_path, "Reports"))# 输出汇总信息print("\n=== RNN建模分析完成 ===")print(f"分析疾病数量: {len(target_diseases)}")print("分析时间范围: 1981-2015(训练) -> 2016-2025(预测)")print(f"使用的气象特征: {', '.join(weather_features)}")print(f"总生成图表数量: {len(target_diseases) * 6 + 2} 个\n")print("主要结果目录:")print("1. Training_History - 训练过程可视化")print("2. Prediction_Comparison - 预测结果对比")print("3. Residual_Analysis - 残差分析")print("4. Performance_Comparison - 模型性能比较")print("5. Prediction_Intervals - 预测区间")print("6. Annual_Comparison - 年度汇总比较")print("7. Seasonal_Patterns - 季节性模式")print("8. Saved_Models - 保存的模型文件")print("9. Reports - 分析报告\n")print("模型性能摘要:")print(performance_summary.to_string(index=False)) best_model = performance_summary.loc[performance_summary['Test_R2'].idxmax()]print("\n最佳性能模型:")print(best_model.to_string())print(f"\n所有结果已保存到: {results_path}")if __name__ == "__main__": main()