# ==================== Python代码:生成纵向数据 ====================import pandas as pdimport numpy as npfrom datetime import datetime, timedeltaimport osfrom pathlib import Pathfrom scipy import statsimport matplotlib.pyplot as pltimport seaborn as snsfrom sklearn.preprocessing import LabelEncoder# 设置随机种子np.random.seed(12345)# 创建结果目录desktop_path = Path.home() / "Desktop"results_dir = desktop_path / "Results"results_dir.mkdir(exist_ok=True)# ==================== 生成模拟纵向数据 ====================def generate_longitudinal_data(n_individuals=100, n_timepoints=5, n_schools=20):""" 生成模拟纵向数据 参数: n_individuals: 个体数量 n_timepoints: 每个个体的时间点数量 n_schools: 学校数量 """
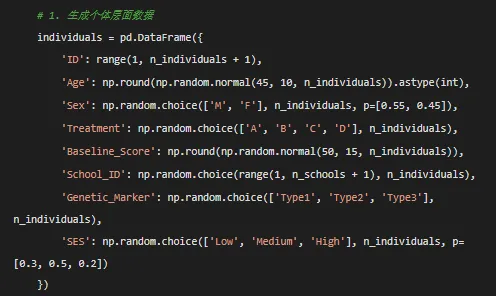
# 2. 生成纵向数据 all_data = []for idx, ind in individuals.iterrows():# 个体特异性随机效应 random_intercept = np.random.normal(0, 2) random_slope = np.random.normal(0, 0.5)# 治疗效应 treatment_map = {'A': 2, 'B': 3, 'C': 1.5, 'D': 0.5} treatment_effect = treatment_map[ind['Treatment']]# 时间序列 time_seq = np.arange(n_timepoints)# 非线性时间效应 time_effect = 0.5 * time_seq + 0.1 * time_seq ** 2 + 0.05 * np.exp(0.2 * time_seq)# 交互效应 age_time_interaction = 0.01 * ind['Age'] * time_seq# 生成连续结局变量 outcome_continuous = ( 30 + random_intercept + (1.5 + random_slope) * time_seq + treatment_effect * time_seq + 0.2 * ind['Age'] + (1.2 if ind['Sex'] == 'M'else 0) + 0.3 * ind['Baseline_Score'] + time_effect + age_time_interaction + np.random.normal(0, 1.5, n_timepoints) )# 二分类结局 logit_prob = 1 / (1 + np.exp(-(0.1 * outcome_continuous - 2))) outcome_binary = np.random.binomial(1, logit_prob)# 计数结局 outcome_count = np.random.poisson(np.exp(0.05 * outcome_continuous + 2))# 生存数据 outcome_survival = np.random.exponential(100, n_timepoints) event_status = np.random.binomial(1, 0.3, n_timepoints)# 多个指标(用于因子分析) indicator1 = 0.7 * outcome_continuous + np.random.normal(0, 2, n_timepoints) indicator2 = 0.6 * outcome_continuous + np.random.normal(0, 2, n_timepoints) indicator3 = 0.5 * outcome_continuous + np.random.normal(0, 2, n_timepoints)# 第二个变量(用于交叉滞后) variable2 = np.zeros(n_timepoints) variable2[0] = 0.6 * outcome_continuous[0] + np.random.normal(0, 1)for t in range(1, n_timepoints): variable2[t] = (0.6 * outcome_continuous[t] + 0.4 * outcome_continuous[t - 1] + np.random.normal(0, 1))# 时变协变量 medication_adherence = np.random.uniform(0.5, 1, n_timepoints) stress_level = np.random.randint(1, 11, n_timepoints) quality_of_life = np.random.uniform(20, 100, n_timepoints)# 测量日期 start_date = datetime(2023, 1, 1) measurement_dates = [start_date + timedelta(days=30 * t) for t in range(n_timepoints)]# 潜在类别 latent_class = np.random.choice([1, 2, 3], p=[0.4, 0.35, 0.25])# 创建个体数据 individual_data = pd.DataFrame({'ID': np.full(n_timepoints, ind['ID']),'Time': time_seq,'Outcome_Continuous': np.round(outcome_continuous, 2),'Outcome_Binary': outcome_binary,'Outcome_Count': outcome_count,'Survival_Time': np.round(outcome_survival, 2),'Event_Status': event_status,'Variable2': np.round(variable2, 2),'Indicator1': np.round(indicator1, 2),'Indicator2': np.round(indicator2, 2),'Indicator3': np.round(indicator3, 2),'Medication_Adherence': np.round(medication_adherence, 2),'Stress_Level': stress_level,'Quality_of_Life': np.round(quality_of_life),'Measurement_Date': measurement_dates,'Age': np.full(n_timepoints, ind['Age']),'Sex': np.full(n_timepoints, ind['Sex']),'Treatment': np.full(n_timepoints, ind['Treatment']),'Baseline_Score': np.full(n_timepoints, ind['Baseline_Score']),'School_ID': np.full(n_timepoints, ind['School_ID']),'Genetic_Marker': np.full(n_timepoints, ind['Genetic_Marker']),'SES': np.full(n_timepoints, ind['SES']),'Latent_Class': np.full(n_timepoints, latent_class),'Random_Intercept': np.full(n_timepoints, random_intercept),'Random_Slope': np.full(n_timepoints, random_slope) }) all_data.append(individual_data)# 合并所有数据 full_data = pd.concat(all_data, ignore_index=True)# 添加缺失值 missing_cols = ['Outcome_Continuous', 'Indicator1', 'Medication_Adherence']for col in missing_cols: mask = np.random.random(len(full_data)) < 0.05 full_data.loc[mask, col] = np.nan# 添加离群值 outlier_idx = np.random.choice(len(full_data), 3, replace=False) full_data.loc[outlier_idx, 'Outcome_Continuous'] *= 2.5return full_data# 生成数据print("正在生成模拟纵向数据...")full_data = generate_longitudinal_data(n_individuals=100, n_timepoints=5)# 创建简化版本simple_data = full_data[['ID', 'Time', 'Outcome_Continuous', 'Treatment', 'Age','Sex', 'Baseline_Score', 'School_ID', 'Measurement_Date']].copy()simple_data = simple_data.rename(columns={'Outcome_Continuous': 'Outcome'})# ==================== 保存数据 ====================# 保存为CSV文件full_data.to_csv(results_dir / "longitudinal_data_full.csv", index=False)simple_data.to_csv(results_dir / "longitudinal_data_simple.csv", index=False)# 保存为Excel文件with pd.ExcelWriter(results_dir / "longitudinal_data.xlsx") as writer: full_data.to_excel(writer, sheet_name='Full_Dataset', index=False) simple_data.to_excel(writer, sheet_name='Simple_Dataset', index=False)# 数据字典 data_dict = pd.DataFrame({'Variable_Name': full_data.columns,'Data_Type': [str(dtype) for dtype in full_data.dtypes],'Description': ['个体唯一标识符','测量时间点(0=基线,1=第1次随访,...)','主要连续结局变量(正态分布)','二分类结局变量(0/1)','计数结局变量(泊松分布)','生存时间(指数分布)','事件状态(0=删失,1=事件)','第二个相关变量(用于交叉滞后分析)','测量指标1(用于因子分析)','测量指标2(用于因子分析)','测量指标3(用于因子分析)','用药依从性比例(0-1)','压力水平评分(1-10)','生活质量评分(20-100)','测量日期','基线年龄','性别(M=男性,F=女性)','治疗分组(A,B,C,D)','基线得分','学校ID(用于多层模型)','遗传标记类型','社会经济状态','潜在类别(模拟不同轨迹组)','随机截距(用于模拟)','随机斜率(用于模拟)' ] }) data_dict.to_excel(writer, sheet_name='Data_Dictionary', index=False)# 保存为Parquet格式(更高效)full_data.to_parquet(results_dir / "longitudinal_data.parquet")simple_data.to_parquet(results_dir / "longitudinal_data_simple.parquet")# ==================== 数据质量检查 ====================print("\n数据质量检查:")print("=" * 50)print(f"总样本量: {len(full_data)} 行")print(f"个体数量: {full_data['ID'].nunique()}")print(f"时间点数量: {full_data['Time'].nunique()}")print(f"缺失值数量: {full_data.isna().sum().sum()}")print("\n连续变量统计摘要:")print(full_data.describe())print("\n类别变量分布:")print(full_data[['Sex', 'Treatment', 'SES']].value_counts().head())# ==================== 可视化数据 ====================print("\n生成可视化图表...")plt.figure(figsize=(15, 12))# 1. 个体轨迹图plt.subplot(2, 2, 1)for i in range(1, 21): # 显示前20个个体的轨迹 individual_data = full_data[full_data['ID'] == i] plt.plot(individual_data['Time'], individual_data['Outcome_Continuous'], alpha=0.5, linewidth=0.8)plt.xlabel('Time')plt.ylabel('Outcome (Continuous)')plt.title('Individual Trajectories')# 2. 治疗组平均轨迹plt.subplot(2, 2, 2)for treatment in ['A', 'B', 'C', 'D']: treatment_data = full_data[full_data['Treatment'] == treatment] mean_trajectory = treatment_data.groupby('Time')['Outcome_Continuous'].mean() plt.plot(mean_trajectory.index, mean_trajectory.values, label=f'Treatment {treatment}', linewidth=2)plt.xlabel('Time')plt.ylabel('Mean Outcome')plt.title('Average Trajectory by Treatment Group')plt.legend()# 3. 潜类别可视化plt.subplot(2, 2, 3)for latent_class in [1, 2, 3]: class_data = full_data[full_data['Latent_Class'] == latent_class] mean_trajectory = class_data.groupby('Time')['Outcome_Continuous'].mean() plt.plot(mean_trajectory.index, mean_trajectory.values, label=f'Latent Class {latent_class}', linewidth=2)plt.xlabel('Time')plt.ylabel('Mean Outcome')plt.title('Trajectory by Latent Class')plt.legend()# 4. 分布直方图plt.subplot(2, 2, 4)plt.hist(full_data['Outcome_Continuous'].dropna(), bins=30, edgecolor='black', alpha=0.7)plt.xlabel('Outcome (Continuous)')plt.ylabel('Frequency')plt.title('Distribution of Outcome Variable')plt.tight_layout()plt.savefig(results_dir / "data_visualization.png", dpi=300, bbox_inches='tight')plt.close()# ==================== 生成Python分析示例代码 ====================python_example_code = '''# ==================== Python纵向数据分析示例 ====================import pandas as pdimport numpy as npimport statsmodels.api as smimport statsmodels.formula.api as smffrom linearmodels.panel import PanelOLSfrom sklearn.mixture import GaussianMixtureimport xgboost as xgbfrom scipy import statsimport matplotlib.pyplot as pltimport seaborn as sns# 1. 混合效应模型import statsmodels.formula.api as smfmodel_mixed = smf.mixedlm("Outcome_Continuous ~ Time * Treatment", full_data, groups=full_data["ID"], re_formula="~Time").fit()print(model_mixed.summary())# 2. 固定效应模型from linearmodels.panel import PanelOLSfull_data_panel = full_data.set_index(['ID', 'Time'])model_fe = PanelOLS.from_formula('Outcome_Continuous ~ Treatment + EntityEffects', data=full_data_panel).fit()# 3. 多层线性模型model_hlm = smf.mixedlm("Outcome_Continuous ~ Time", full_data, groups=full_data["School_ID"], re_formula="~Time").fit()# 4. 广义估计方程model_gee = smf.gee("Outcome_Binary ~ Time * Treatment", "ID", full_data, cov_struct=sm.cov_struct.Exchangeable(), family=sm.families.Binomial()).fit()# 5. 广义线性混合模型 (GLMM)# 可以使用statsmodels或专门的包# 9. 重复测量ANOVAimport pingouin as pgrm_anova = pg.rm_anova(data=full_data, dv='Outcome_Continuous', within='Time', subject='ID')# 13. 广义相加模型 (GAM)from pygam import LinearGAM, smodel_gam = LinearGAM(s(0) + s(1)).fit(full_data[['Time', 'Age']], full_data['Outcome_Continuous'])# 14-15. 潜类别/剖面模型from sklearn.mixture import GaussianMixturegmm = GaussianMixture(n_components=3).fit(full_data[['Outcome_Continuous', 'Stress_Level', 'Quality_of_Life']])full_data['cluster'] = gmm.predict(full_data[['Outcome_Continuous', 'Stress_Level', 'Quality_of_Life']])# 16. 状态空间模型from statsmodels.tsa.statespace.sarimax import SARIMAXmodel_ssm = SARIMAX(full_data['Outcome_Continuous'], order=(1,0,0)).fit()# 18. 贝叶斯模型 (使用pymc3)import pymc3 as pmwith pm.Model(): alpha = pm.Normal('alpha', mu=0, sigma=10) beta = pm.Normal('beta', mu=0, sigma=10) mu = alpha + beta * full_data['Time'] y_obs = pm.Normal('y_obs', mu=mu, sigma=1, observed=full_data['Outcome_Continuous']) trace = pm.sample(1000)# 19. 混合效应随机森林# 可以使用merf库# 20. 纵向梯度提升dtrain = xgb.DMatrix(full_data[['Time', 'Age', 'Baseline_Score']], label=full_data['Outcome_Continuous'])params = {'objective': 'reg:squarederror', 'max_depth': 3}model_xgb = xgb.train(params, dtrain, num_boost_round=100)# 21. K均值纵向聚类# 将数据转换为宽格式wide_data = full_data.pivot(index='ID', columns='Time', values='Outcome_Continuous')from sklearn.cluster import KMeanskmeans = KMeans(n_clusters=3).fit(wide_data.fillna(wide_data.mean()))# 22. 基于模型的聚类from sklearn.mixture import GaussianMixturegmm_cluster = GaussianMixture(n_components=3).fit(full_data[['Outcome_Continuous', 'Time']])full_data['model_cluster'] = gmm_cluster.predict(full_data[['Outcome_Continuous', 'Time']])'''# 保存Python示例代码with open(results_dir / "example_analysis_python.py", "w", encoding="utf-8") as f: f.write(python_example_code)# ==================== 生成元数据文件 ====================metadata = {"dataset_info": {"name": "Longitudinal_Mixed_Effects_Dataset","version": "1.0","creation_date": datetime.now().strftime("%Y-%m-%d"),"description": "Simulated longitudinal dataset for testing various statistical models" },"dimensions": {"n_individuals": int(full_data['ID'].nunique()),"n_timepoints": int(full_data['Time'].nunique()),"n_variables": int(len(full_data.columns)),"total_observations": int(len(full_data)) },"variables": [ {"name": col,"type": str(full_data[col].dtype),"n_unique": int(full_data[col].nunique()) if full_data[col].dtype == 'object'else None,"missing": int(full_data[col].isna().sum()) }for col in full_data.columns ],"generation_parameters": {"random_seed": 12345,"n_individuals": 100,"n_timepoints": 5,"treatment_groups": ["A", "B", "C", "D"],"time_effect": "linear + quadratic + exponential","random_effects": ["intercept", "slope"] }}import jsonwith open(results_dir / "metadata.json", "w", encoding="utf-8") as f: json.dump(metadata, f, indent=2, ensure_ascii=False)# ==================== 输出总结信息 ====================print("\n" + "=" * 60)print("数据生成完成!")print("=" * 60 + "\n")print("生成的文件:")files_generated = ["longitudinal_data.xlsx","longitudinal_data_full.csv","longitudinal_data_simple.csv","longitudinal_data.parquet","longitudinal_data_simple.parquet","data_visualization.png","example_analysis_python.py","metadata.json"]for file in files_generated: file_path = results_dir / fileprint(f" ✓ {file} ({file_path.stat().st_size / 1024:.1f} KB)")print(f"\n数据集结构:")print(f" 总行数: {len(full_data)}")print(f" 总列数: {len(full_data.columns)}")print(f" 个体数: {full_data['ID'].nunique()}")print(f" 时间点: {full_data['Time'].max() + 1}")print("\n变量类型:")dtype_counts = full_data.dtypes.value_counts()for dtype, count in dtype_counts.items():print(f" {dtype}: {count} 个变量")print(f"\n数据已保存到: {results_dir}")print("\n可以使用以下代码加载数据:")print(f" R语言: load('{results_dir / 'longitudinal_data.RData'}')")print(f" Python: pd.read_csv('{results_dir / 'longitudinal_data_full.csv'}')")# 显示前几行数据示例print("\n数据示例(前5行):")print(full_data.head().to_string())