# pip install pandas numpy matplotlib seaborn scikit-learn dtw-python joblibimport osimport pandas as pdimport numpy as npimport matplotlib.pyplot as pltimport seaborn as snsfrom sklearn.cluster import KMeansfrom sklearn.neighbors import KNeighborsClassifierfrom sklearn.metrics import accuracy_scorefrom sklearn.preprocessing import StandardScalerfrom scipy.spatial.distance import pdist, squareformimport dtwfrom datetime import datetimeimport warningswarnings.filterwarnings('ignore')# 设置路径desktop_path = os.path.join(os.path.expanduser("~"), "Desktop")data_path = os.path.join(desktop_path, "Results", "combined_weather_disease_data.csv")results_path = os.path.join(desktop_path, "Results时间EM")# 设置中文字体显示plt.rcParams['font.sans-serif'] = ['SimHei'] # 显示中文标签plt.rcParams['axes.unicode_minus'] = False # 显示负号# 创建结果文件夹sub_folders = ['Clustering', 'KNN', 'DTW', 'Plots', 'Models']for folder in sub_folders: os.makedirs(os.path.join(results_path, folder), exist_ok=True)print("开始基于距离的疾病时间序列分析...")# 读取数据combined_data = pd.read_csv(data_path)combined_data['timestamp'] = pd.to_datetime(combined_data['timestamp'])# 选择要分析的疾病target_diseases = ["influenza", "common_cold", "pneumonia","bacillary_dysentery", "hand_foot_mouth","hemorrhagic_fever"]print(f"目标疾病: {', '.join(target_diseases)}")# 数据预处理函数 - 按年汇总数据def prepare_yearly_data(data, diseases): data = data.copy() data['year'] = data['timestamp'].dt.year yearly_data = data.groupby('year')[diseases].sum().reset_index() yearly_data = yearly_data[(yearly_data['year'] >= 1981) & (yearly_data['year'] <= 2025)]return yearly_data# 数据预处理函数 - 创建时间序列矩阵def create_ts_matrix(data, diseases): ts_matrix = data.set_index('year')[diseases].valuesreturn ts_matrix# 应用数据预处理print("正在准备数据...")yearly_data = prepare_yearly_data(combined_data, target_diseases)ts_matrix = create_ts_matrix(yearly_data, target_diseases)years = yearly_data['year'].values# 划分训练集和测试集train_years = range(1981, 2016)test_years = range(2016, 2026)train_mask = yearly_data['year'].isin(train_years)test_mask = yearly_data['year'].isin(test_years)train_data = yearly_data[train_mask]test_data = yearly_data[test_mask]train_matrix = ts_matrix[train_mask]test_matrix = ts_matrix[test_mask]print(f"训练集大小: {len(train_data)} 年")print(f"测试集大小: {len(test_data)} 年")
# 选择最佳k值(这里选择3)optimal_k = 3# 执行K-Means聚类kmeans = KMeans(n_clusters=optimal_k, random_state=123, n_init=25)kmeans_result = kmeans.fit_predict(ts_matrix)# 添加聚类标签yearly_data['cluster'] = kmeans_resultyearly_data['cluster'] = yearly_data['cluster'].astype(str)# 2. 聚类结果可视化# 2.1 聚类中心图cluster_centers = pd.DataFrame( kmeans.cluster_centers_, columns=target_diseases)cluster_centers['cluster'] = [str(i) for i in range(optimal_k)]cluster_centers_long = pd.melt( cluster_centers, id_vars=['cluster'], var_name='disease', value_name='cases')plt.figure(figsize=(12, 8))sns.barplot(data=cluster_centers_long, x='disease', y='cases', hue='cluster')plt.title('聚类中心 - 各疾病平均病例数')plt.xlabel('疾病类型')plt.ylabel('平均病例数')plt.xticks(rotation=45)plt.legend(title='聚类')plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'Cluster_Centers.png'), dpi=300, bbox_inches='tight')plt.close()# 2.2 聚类分配时间线plt.figure(figsize=(12, 6))for cluster in yearly_data['cluster'].unique(): cluster_data = yearly_data[yearly_data['cluster'] == cluster] plt.scatter(cluster_data['year'], [1] * len(cluster_data), label=f'聚类 {cluster}', s=100)plt.plot(yearly_data['year'], [1] * len(yearly_data), 'gray', alpha=0.5)plt.title('聚类分配时间线 (1981-2025)')plt.xlabel('年份')plt.yticks([])plt.legend(title='聚类')plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'Cluster_Timeline.png'), dpi=300, bbox_inches='tight')plt.close()
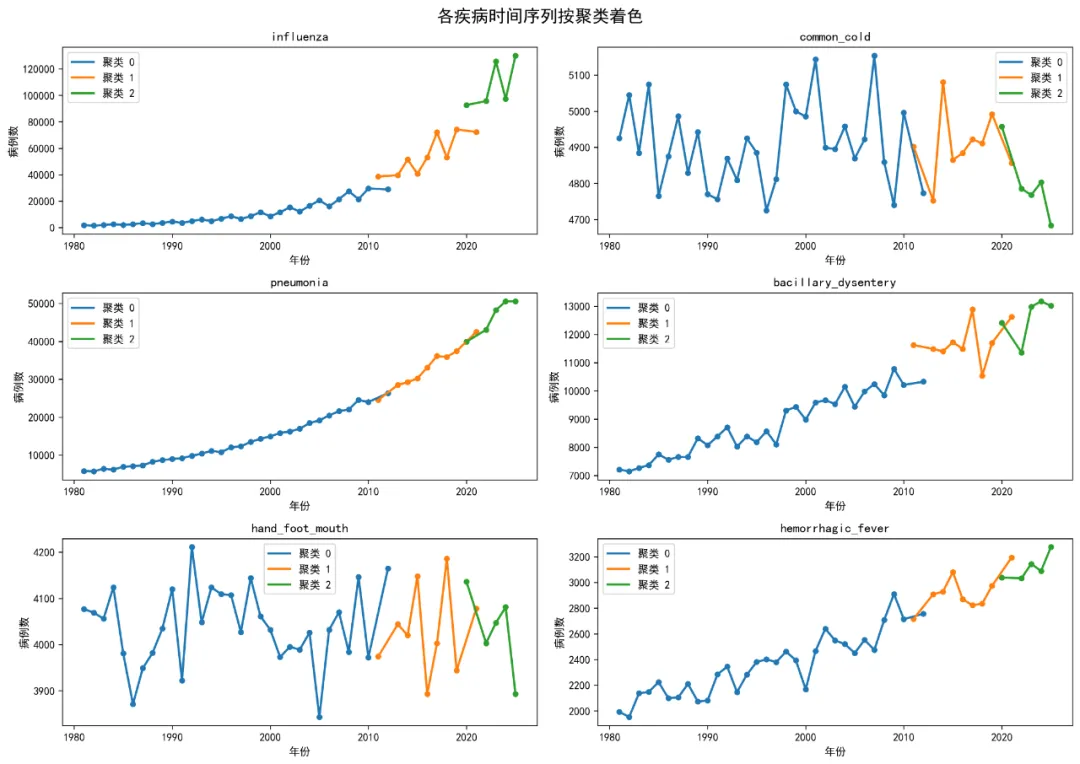
# 2.3 多序列对比图(按聚类着色)yearly_data_long = pd.melt( yearly_data, id_vars=['year', 'cluster'], value_vars=target_diseases, var_name='disease', value_name='cases')fig, axes = plt.subplots(3, 2, figsize=(14, 10))axes = axes.flatten()for i, disease in enumerate(target_diseases): disease_data = yearly_data_long[yearly_data_long['disease'] == disease]for cluster in disease_data['cluster'].unique(): cluster_data = disease_data[disease_data['cluster'] == cluster] axes[i].plot(cluster_data['year'], cluster_data['cases'], label=f'聚类 {cluster}', linewidth=2) axes[i].scatter(cluster_data['year'], cluster_data['cases'], s=20) axes[i].set_title(disease) axes[i].set_xlabel('年份') axes[i].set_ylabel('病例数') axes[i].legend()plt.suptitle('各疾病时间序列按聚类着色', fontsize=16)plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'Multi_Series_Clusters.png'), dpi=300, bbox_inches='tight')plt.close()
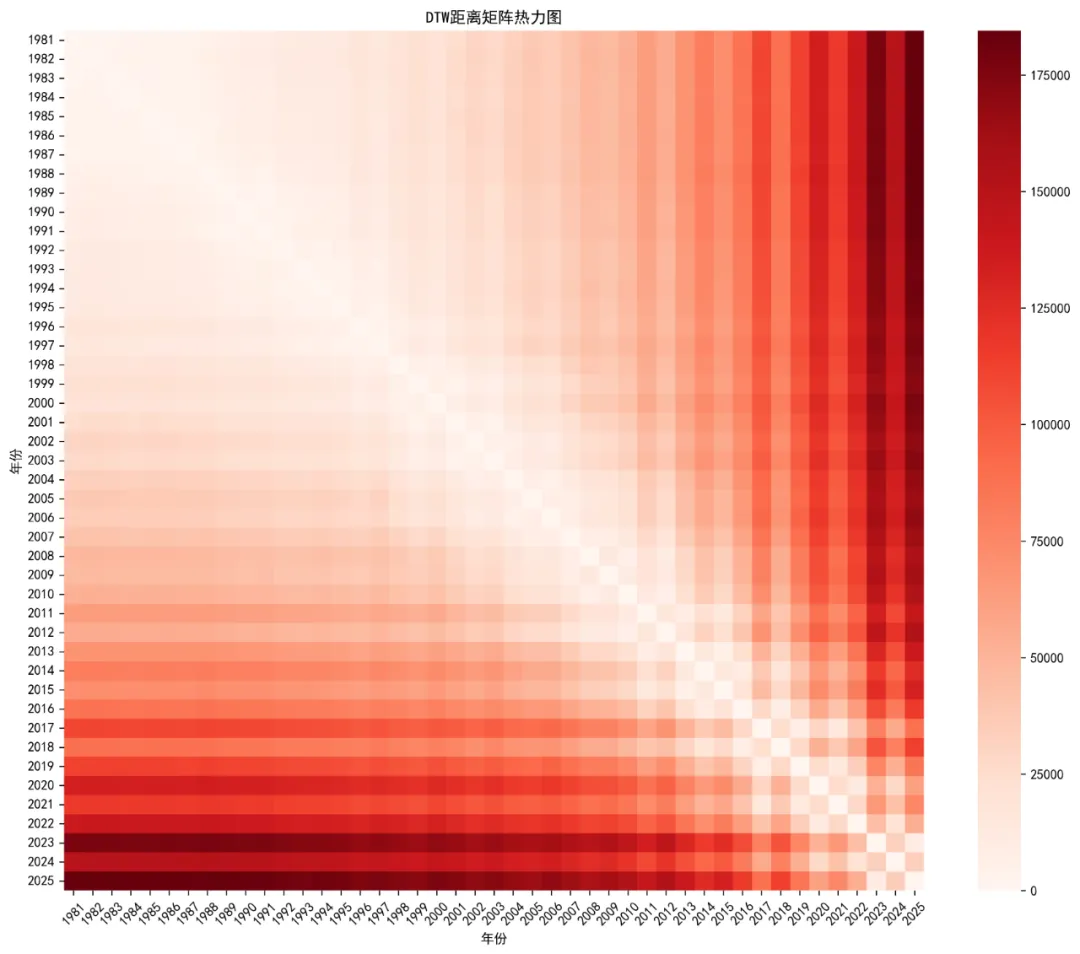
# 2.4 聚类热力图from sklearn.decomposition import PCA# 使用PCA降维进行可视化pca = PCA(n_components=2)X_pca = pca.fit_transform(ts_matrix)plt.figure(figsize=(10, 8))scatter = plt.scatter(X_pca[:, 0], X_pca[:, 1], c=kmeans_result, cmap='viridis', s=100)plt.colorbar(scatter, label='聚类')plt.title('K-Means聚类结果可视化')plt.xlabel('主成分 1')plt.ylabel('主成分 2')# 添加年份标签for i, year in enumerate(years): plt.annotate(str(year), (X_pca[i, 0], X_pca[i, 1]), xytext=(5, 5), textcoords='offset points', fontsize=8)plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'Cluster_Heatmap.png'), dpi=300, bbox_inches='tight')plt.close()# 3. 基于KNN的时间序列分类print("正在进行KNN分类分析...")# 准备KNN数据knn_train = train_matrixknn_test = test_matrixknn_train_labels = kmeans_result[train_mask]# 寻找最佳k值k_values = range(1, 11)accuracy_values = []for k in k_values: knn = KNeighborsClassifier(n_neighbors=k) knn.fit(knn_train, knn_train_labels) knn_pred = knn.predict(knn_test) accuracy = accuracy_score(kmeans_result[test_mask], knn_pred) accuracy_values.append(accuracy)# K值选择图plt.figure(figsize=(10, 6))plt.plot(k_values, accuracy_values, 'bo-', linewidth=2, markersize=8, color='steelblue')plt.xlabel('K值')plt.ylabel('测试集准确率')plt.title('KNN分类准确率 vs K值')plt.xticks(k_values)plt.grid(True, alpha=0.3)plt.savefig(os.path.join(results_path, 'Plots', 'K_Selection.png'), dpi=300, bbox_inches='tight')plt.close()# 选择最佳k值best_k = k_values[np.argmax(accuracy_values)]print(f"最佳K值: {best_k}, 准确率: {max(accuracy_values):.3f}")# 使用最佳k值进行KNN预测knn = KNeighborsClassifier(n_neighbors=best_k)knn.fit(knn_train, knn_train_labels)knn_proba = knn.predict_proba(knn_test)knn_pred = knn.predict(knn_test)# KNN预测结果knn_results = pd.DataFrame({'year': test_data['year'].values,'actual_cluster': kmeans_result[test_mask].astype(str),'predicted_cluster': knn_pred.astype(str),'probability': np.max(knn_proba, axis=1),'correct': (knn_pred == kmeans_result[test_mask])})# 4. DTW(动态时间规整)分析print("正在进行DTW分析...")# 计算DTW距离矩阵n_years = len(ts_matrix)dtw_dist = np.zeros((n_years, n_years))# 使用更高效的DTW计算for i in range(n_years):for j in range(i, n_years):if i == j: dtw_dist[i, j] = 0else: distance = dtw.dtw(ts_matrix[i], ts_matrix[j]).distance dtw_dist[i, j] = distance dtw_dist[j, i] = distance# DTW距离热力图plt.figure(figsize=(12, 10))sns.heatmap(dtw_dist, cmap='Reds', xticklabels=years, yticklabels=years)plt.title('DTW距离矩阵热力图')plt.xlabel('年份')plt.ylabel('年份')plt.xticks(rotation=45)plt.yticks(rotation=0)plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'DTW_Heatmap.png'), dpi=300, bbox_inches='tight')plt.close()# 查找最具代表性的序列(距离其他序列平均距离最小的)avg_dtw_dist = np.mean(dtw_dist, axis=1)most_typical_idx = np.argmin(avg_dtw_dist)most_atypical_idx = np.argmax(avg_dtw_dist)most_typical_year = years[most_typical_idx]most_atypical_year = years[most_atypical_idx]print(f"最具代表性的年份: {most_typical_year}")print(f"最不典型的年份: {most_atypical_year}")# 5. 公共卫生意义分析 - 识别疫情模式print("正在进行公共卫生模式分析...")# 分析每个聚类的特征cluster_profiles = yearly_data.groupby('cluster')[target_diseases].agg(['mean', 'std'])# 重命名列cluster_profiles.columns = [f'{col[0]}_{col[1]}'for col in cluster_profiles.columns]cluster_profiles = cluster_profiles.reset_index()# 识别聚类模式cluster_patterns = pd.DataFrame({'cluster': [str(i) for i in range(optimal_k)],'pattern_type': ['快速爆发型', '缓慢持续型', '平稳低发型'][:optimal_k],'description': ['病例数快速上升下降', '病例数持续较高水平', '病例数保持较低水平'][:optimal_k]})# 6. 生成综合报告图表# 7.1 聚类性能总结图fig, axes = plt.subplots(2, 2, figsize=(16, 12))# 肘部法则图axes[0,0].plot(k_range, wss, 'bo-', linewidth=2, markersize=8, color='steelblue')axes[0,0].set_xlabel('聚类数量 (k)')axes[0,0].set_ylabel('组内平方和')axes[0,0].set_title('K-Means聚类肘部法则')axes[0,0].set_xticks(k_range)axes[0,0].grid(True, alpha=0.3)# 聚类中心图sns.barplot(data=cluster_centers_long, x='disease', y='cases', hue='cluster', ax=axes[0,1])axes[0,1].set_title('聚类中心 - 各疾病平均病例数')axes[0,1].set_xlabel('疾病类型')axes[0,1].set_ylabel('平均病例数')axes[0,1].tick_params(axis='x', rotation=45)# 聚类分配时间线for cluster in yearly_data['cluster'].unique(): cluster_data = yearly_data[yearly_data['cluster'] == cluster] axes[1,0].scatter(cluster_data['year'], [1] * len(cluster_data), label=f'聚类 {cluster}', s=100)axes[1,0].plot(yearly_data['year'], [1] * len(yearly_data), 'gray', alpha=0.5)axes[1,0].set_title('聚类分配时间线 (1981-2025)')axes[1,0].set_xlabel('年份')axes[1,0].set_yticks([])axes[1,0].legend(title='聚类')# 聚类热力图scatter = axes[1,1].scatter(X_pca[:, 0], X_pca[:, 1], c=kmeans_result, cmap='viridis', s=100)axes[1,1].set_title('K-Means聚类结果可视化')axes[1,1].set_xlabel('主成分 1')axes[1,1].set_ylabel('主成分 2')plt.suptitle('基于欧氏距离的聚类分析总结', fontsize=16)plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'Cluster_Analysis_Summary.png'), dpi=300, bbox_inches='tight')plt.close()# 7.2 KNN预测结果图plt.figure(figsize=(12, 8))colors = ['red'if not correct else'green'for correct in knn_results['correct']]plt.scatter(knn_results['year'], knn_results['correct'].astype(int), c=colors, s=200, alpha=0.7)for _, row in knn_results.iterrows(): plt.annotate(f"实际:{row['actual_cluster']}\n预测:{row['predicted_cluster']}", (row['year'], row['correct']), xytext=(5, 5), textcoords='offset points', fontsize=9)accuracy_rate = knn_results['correct'].mean() * 100plt.title(f'KNN分类预测结果 (k = {best_k})\n准确率: {accuracy_rate:.1f}%')plt.xlabel('年份')plt.ylabel('分类正确性')plt.yticks([0, 1], ['错误', '正确'])plt.grid(True, alpha=0.3)plt.tight_layout()plt.savefig(os.path.join(results_path, 'Plots', 'KNN_Prediction_Results.png'), dpi=300, bbox_inches='tight')plt.close()# 8. 保存数据结果# 保存聚类结果yearly_data.to_csv(os.path.join(results_path, 'Clustering', 'Yearly_Data_with_Clusters.csv'), index=False)cluster_centers.to_csv(os.path.join(results_path, 'Clustering', 'Cluster_Centers.csv'), index=False)cluster_profiles.to_csv(os.path.join(results_path, 'Clustering', 'Cluster_Profiles.csv'), index=False)cluster_patterns.to_csv(os.path.join(results_path, 'Clustering', 'Cluster_Patterns.csv'), index=False)# 保存KNN结果knn_results.to_csv(os.path.join(results_path, 'KNN', 'KNN_Prediction_Results.csv'), index=False)# 保存DTW结果dtw_df = pd.DataFrame(dtw_dist, index=years, columns=years)dtw_df.to_csv(os.path.join(results_path, 'DTW', 'DTW_Distance_Matrix.csv'))# 保存模型import joblibjoblib.dump(kmeans, os.path.join(results_path, 'Models', 'kmeans_model.pkl'))joblib.dump(knn, os.path.join(results_path, 'Models', 'knn_model.pkl'))# 9. 生成分析报告print("\n=== 基于距离的时间序列分析完成 ===")print(f"分析方法: 欧氏距离 + K-Means聚类 + KNN分类 + DTW")print(f"分析疾病: {', '.join(target_diseases)}")print(f"时间范围: 1981-2025")print(f"训练集: 1981-2015 ({len(train_data)} 年)")print(f"测试集: 2016-2025 ({len(test_data)} 年)\n")print("聚类分析结果:")print(f"最佳聚类数量: {optimal_k}")print("聚类分布:")print(yearly_data['cluster'].value_counts().sort_index())print("\nKNN分类结果:")print(f"最佳K值: {best_k}")print(f"测试集准确率: {accuracy_rate:.1f}%")print("\nDTW分析结果:")print(f"最具代表性年份: {most_typical_year}")print(f"最不典型年份: {most_atypical_year}")print("\n公共卫生模式识别:")print(cluster_patterns.to_string(index=False))print("\n文件输出:")print("1. Results时间EM/Clustering/ - 聚类分析结果")print("2. Results时间EM/KNN/ - KNN分类结果")print("3. Results时间EM/DTW/ - DTW分析结果")print("4. Results时间EM/Plots/ - 所有可视化图表")print("5. Results时间EM/Models/ - 训练好的模型")print("\n公共卫生意义:")print("• 快速爆发型: 需要建立快速响应机制和应急预案")print("• 缓慢持续型: 需要长期监测和持续防控措施")print("• 平稳低发型: 可作为基线参考,重点关注异常波动")# 性能分析performance_summary = pd.DataFrame({'指标': ['聚类数量', 'KNN最佳K值', 'KNN准确率', '最具代表性年份', '最不典型年份'],'数值': [optimal_k, best_k, f'{accuracy_rate:.1f}%', most_typical_year, most_atypical_year]})performance_summary.to_csv(os.path.join(results_path, 'Performance_Summary.csv'), index=False)print("\n=== 分析完成 ===")