NG文章的空间转录组没给h5怎么办:用Python手动构建AnnData,让空间坐标、图片、尺度信息都对上
- 2026-02-27 00:16:38
NG文章的空间转录组没给h5怎么办:用Python手动构建AnnData,让空间坐标、图片、尺度信息都对上


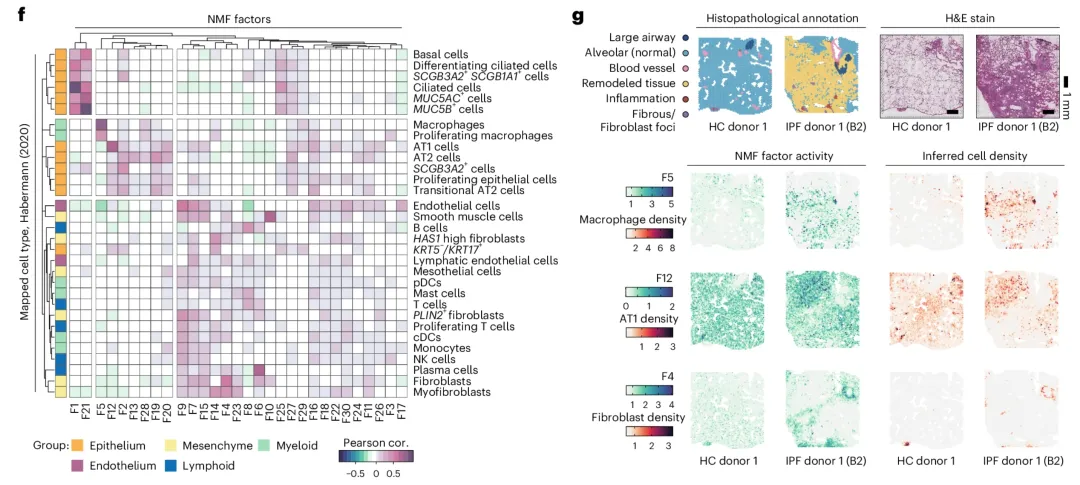
文章数据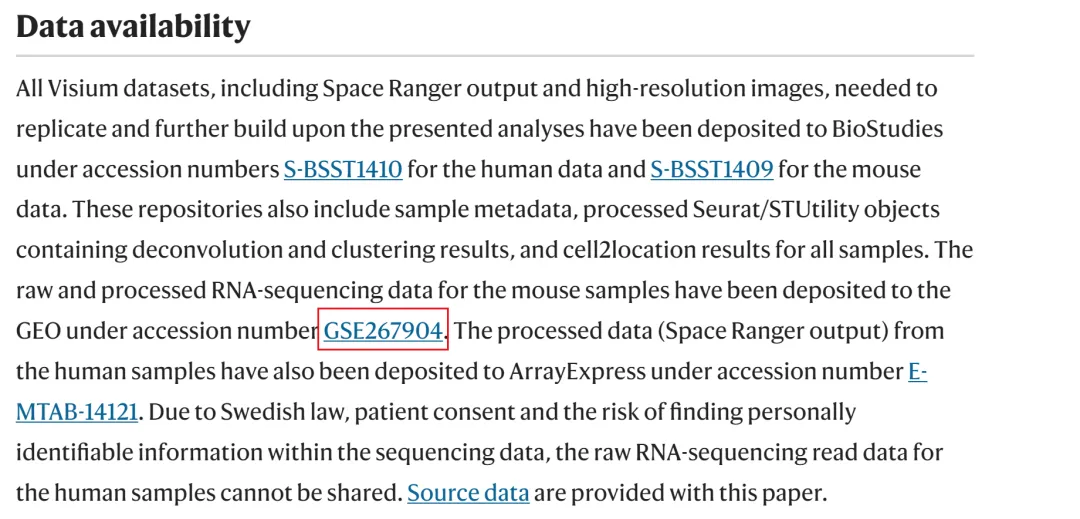 
在GEO中下载数据 
文章一共测了24个样本,点击红色框中的 custom 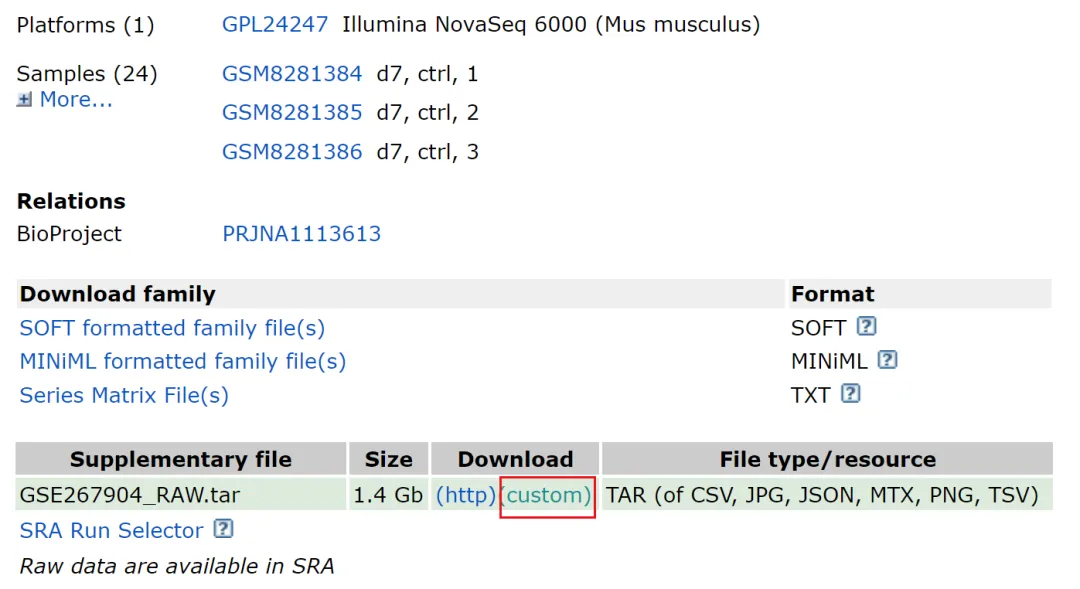
下载第一个样本,样本一共包含有九个文件 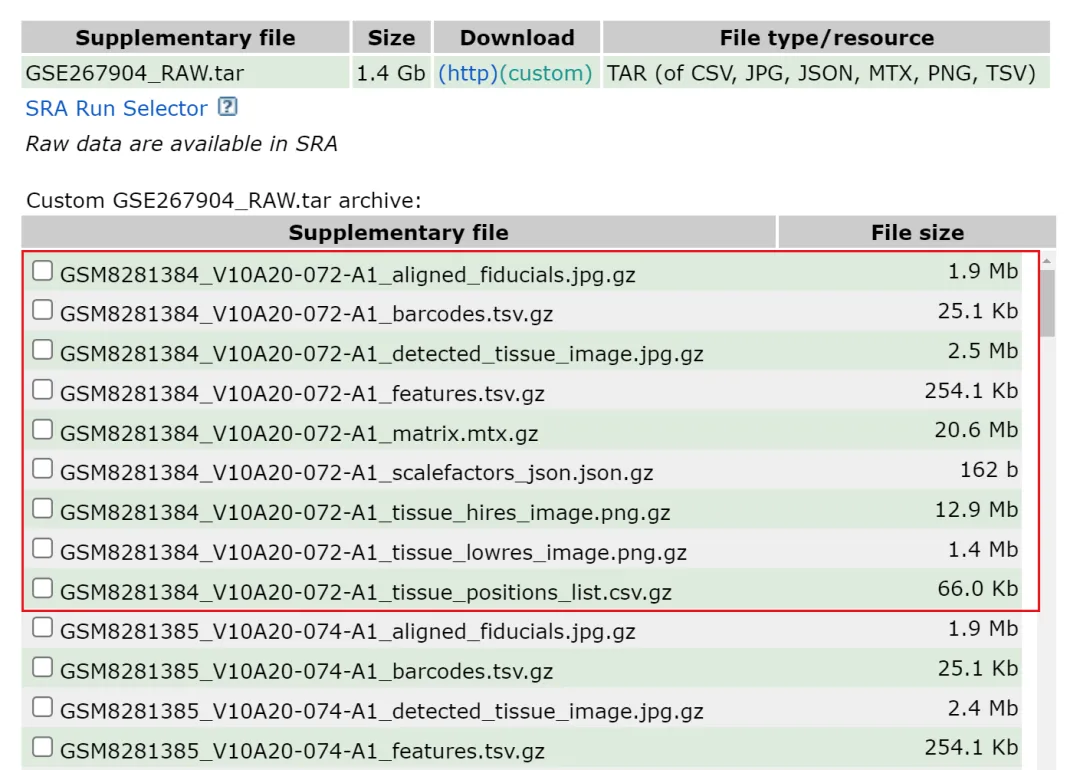
文件处理---主要是修改文件名字 1.矩阵文件名字必须改为barcodes.tsv.gz features.tsv.gz和matrix.mtx.gz 三个标准的文件名字,建议把三个文件放到一个文件夹中 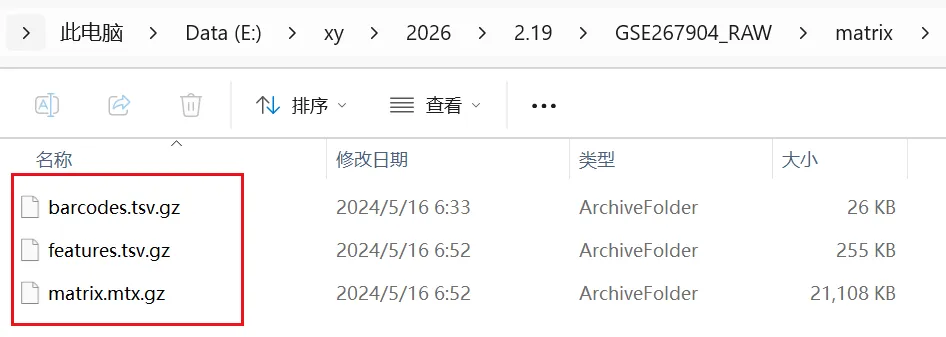
2)读取 图像缩放比例,用于把 fullres 坐标映射到 hires/lowres 图上 3)读取 3.1 文件结构 3.2 自动处理 “-1” 后缀对齐(处理有没有 -1 的情况) 4)写入 5)加载 hires/lowres 图片 6)写入 7)快速检查 + 第一张 spatial 图 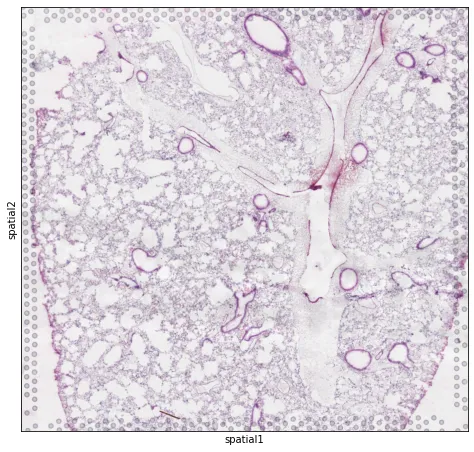
8)空转数据标准分析流程 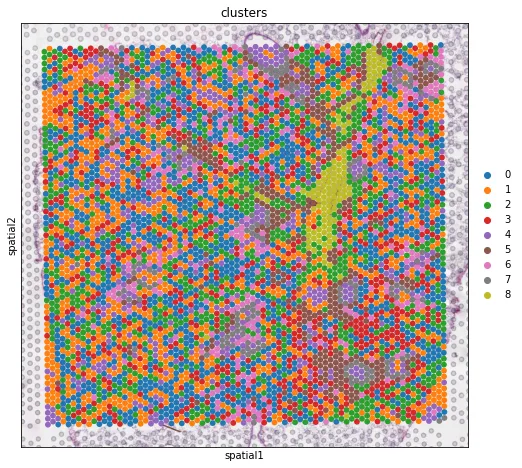
数据保存 零基础也能学的AI生信课(AI助力生信入门班即将开始) AI助力多组学与机器学习联合分析(机器学习分析代谢组、蛋白组、宏基因组、网络药理学、转录组) CNS文章单细胞数据分析内容汇总(基础--进阶--python分析) Nature单细胞空间组学1:1复现训练营 跟着CNS文章学空间转录组分析线上直播课(Visium HD、Xenium、Stereo-seq) 




NG | 文献







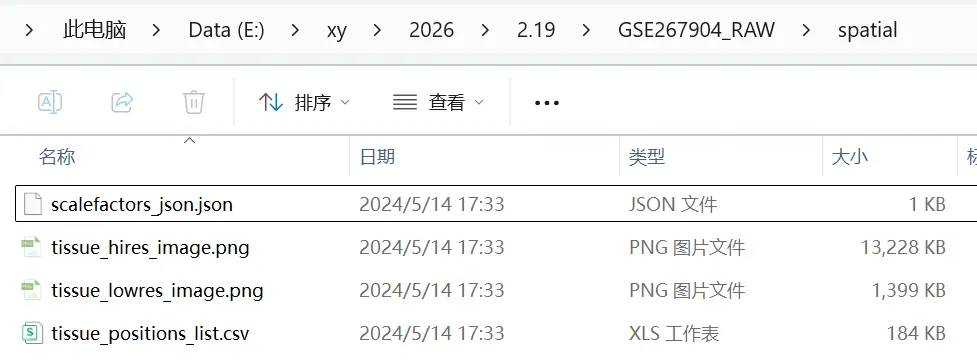
2.创建一个 spatial 的文件夹,文件夹中必须包含下面四个文件

分析流程
0)导入库 + 文件路径
import scanpy as scimport matplotlib.pyplot as pltimport seaborn as snsimport gzip, ioimport numpy as np, pandas as pdimport scipy.io, scipy.sparse as spimport h5pyimport os, jsonfrom PIL import Image
设置路径
MTX_DIR = r"E:\xy\2026\2.19\GSE267904_RAW\\matrix\\" # 含 barcodes.tsv.gz / features.tsv.gz / matrix.mtx.gzSPATIAL_DIR = r"E:\xy\2026\2.19\GSE267904_RAW\\spatial" # 含 tissue_positions_list.csv / scalefactors_json.json / tissue_*_image.pngLIBRARY_ID = "072-A1"
1)读取 10x 表达矩阵
adata = sc.read_10x_mtx(MTX_DIR,var_names="gene_symbols", # 你也可以换成 gene_idscache=True)adata.var_names_make_unique()
scalefactors_json.jsonsf_path = os.path.join(SPATIAL_DIR,"scalefactors_json.json")withopen(sf_path, "r", encoding="utf-8") as f:scalefactors = json.load(f)
tissue_positions_list.csv 并对齐 barcodepos_path = os.path.join(SPATIAL_DIR, "tissue_positions_list.csv")pos = pd.read_csv(pos_path, header=None)# 10x Visium 标准(无表头)6列:# barcode, in_tissue, array_row, array_col, pxl_row_in_fullres, pxl_col_in_fullrespos.columns = ["barcode", "in_tissue", "array_row","array_col", "pxl_row_in_fullres","pxl_col_in_fullres"]pos = pos.set_index("barcode")pos.index.name = None
obs_names = adata.obs_names.astype(str)if obs_names[0].endswith("-1") and (not str(pos.index[0]).endswith("-1")):pos.index = pos.index.astype(str) + "-1"elif (not obs_names[0].endswith("-1")) and str(pos.index[0]).endswith("-1"):pos.index = pos.index.astype(str).str.replace("-1$", "", regex=True)# 只保留 adata 里有的条码顺序pos = pos.reindex(obs_names)
obs 和 obsm["spatial"]adata.obs = adata.obs.join(pos[["in_tissue", "array_row","array_col", "pxl_row_in_fullres","pxl_col_in_fullres"]])# 重点:obsm["spatial"] 放 fullres 坐标;不要乘 tissue_hires_scalef / lowres_scalef# 坐标顺序:x=col, y=row(更符合图像坐标:横轴列、纵轴行)adata.obsm["spatial"] = pos[["pxl_col_in_fullres","pxl_row_in_fullres"]].to_numpy()
img_candidates = [("hires", os.path.join(SPATIAL_DIR, "tissue_hires_image.png")),("lowres", os.path.join(SPATIAL_DIR, "tissue_lowres_image.png")),]images = {}for k, p in img_candidates:if os.path.exists(p):im = Image.open(p).convert("RGB")images[k] = (np.asarray(im) / 255.0).astype(np.float32)iflen(images) == 0:print("WARNING: 没找到 tissue_hires_image.png 或 tissue_lowres_image.png,后续 sc.pl.spatial 只能画点图不叠图。")
uns["spatial"](Scanpy 识别 Visium 的标准结构)adata.uns["spatial"] = {LIBRARY_ID: {}}adata.uns["spatial"][LIBRARY_ID]["scalefactors"] = scalefactorsadata.uns["spatial"][LIBRARY_ID]["images"] = images# 如果同时有 hires/lowres,默认用 hires;没有就用 lowresif "hires" in images:adata.uns["spatial"][LIBRARY_ID]["use_quality"] = "hires"elif "lowres" in images:adata.uns["spatial"][LIBRARY_ID]["use_quality"] = "lowres"
print(adata)print("scalefactors:", adata.uns["spatial"][LIBRARY_ID]["scalefactors"])# 叠加组织图(优先 hires)img_key = "hires" if "hires" in images else ("lowres" if "lowres" in images else None)if img_key is not None:sc.pl.spatial(adata, library_id=LIBRARY_ID, img_key=img_key, spot_size=1.2)else:# 没有图片也能画(只是没有底图)sc.pl.embedding(adata, basis="spatial")

# 标记线粒体基因adata.var["mt"] = adata.var_names.str.startswith("Mt-")sc.pp.calculate_qc_metrics(adata, qc_vars=["mt"], inplace=True)print(f"#cells after MT filter: {adata.n_obs}")sc.pp.filter_genes(adata, min_cells=10)sc.pp.normalize_total(adata, inplace=True)sc.pp.log1p(adata)sc.pp.highly_variable_genes(adata, flavor="seurat", n_top_genes=2000)sc.pp.pca(adata)sc.pp.neighbors(adata)sc.tl.umap(adata)sc.tl.leiden(adata, key_added="clusters" )plt.rcParams["figure.figsize"] = (4, 4)sc.pl.umap(adata,color=["total_counts", "n_genes_by_counts", "clusters"],wspace=0.4)sc.pl.umap(adata, color=["total_counts" ], wspace=0.4)sc.pl.umap(adata, color=[ "n_genes_by_counts"], wspace=0.4)sc.pl.umap(adata, color=[ "clusters"], wspace=0.4)plt.rcParams["figure.figsize"] = (8, 8)sc.pl.spatial(adata, img_key="hires",color=["total_counts", "n_genes_by_counts"])sc.pl.spatial(adata, img_key="hires",color="clusters", size=1.5)sc.tl.rank_genes_groups(adata, "clusters", method="t-test")sc.pl.rank_genes_groups_heatmap(adata, groups="2", n_genes=10, groupby="clusters")sc.pl.spatial(adata, img_key="hires",color=["clusters" ])

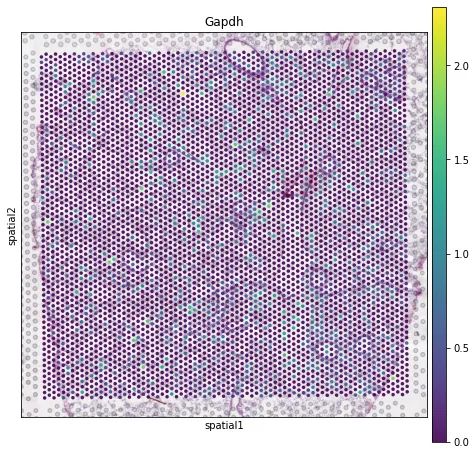
sc.pl.spatial(adata, img_key="hires",color=["Gapdh"], alpha=0.9)

adata.write_h5ad(r"E:\xy\2026\2.19\A1_visium_fixed.h5ad")零基础想学生信,看👇这个文章:
想学机器学习,看👇这个文章:
想系统学单细胞,看👇这个文章:
想系统学空间转录组,看👇这个文章:
欢迎关注 | 华哥科研平台


亲,写的这么辛苦,记得关注、点赞、转发哟!

本文来自网友投稿或网络内容,如有侵犯您的权益请联系我们删除,联系邮箱:wyl860211@qq.com 。
